Show code
library(brms)
library(tidyverse)
library(bayesplot)
# Set seed for reproducibility
set.seed(42)Bayesian Mixed Effects Models with brms for Linguists
After fitting your model, validate that it generates data similar to what you observed.
Posterior predictive checks answer: “If I were to generate new data from my fitted model, would it look like my actual data?” This validates that your model has captured the essential structure of your data.
For binary outcomes (correct/incorrect, yes/no), we focus on: - Proportion of successes (mean of 0/1 outcomes) - Accuracy by condition - Calibration (predicted probabilities match observed frequencies)
library(brms)
library(tidyverse)
library(bayesplot)
# Set seed for reproducibility
set.seed(42)# Create example grammaticality judgment data (same as prior checks)
n_subj <- 25
n_trials <- 40
n_items <- 30
gram_data <- expand.grid(
trial = 1:n_trials,
subject = 1:n_subj,
item = 1:n_items
) %>%
filter(row_number() <= n_subj * n_trials * 2) %>%
mutate(
condition = rep(c('A', 'B'), length.out = n()),
p_correct = plogis(0.2 + (condition == 'B') * 0.4),
correct = rbinom(n(), size = 1, prob = p_correct)
)
# Define priors
gram_priors <- c(
prior(normal(0, 1.5), class = Intercept),
prior(normal(0, 1), class = b),
prior(exponential(1), class = sd),
prior(lkj(2), class = cor)
)
# Check if model exists, otherwise fit it
model_file <- "fits/fit_gram.rds"
if (file.exists(model_file)) {
cat("Loading saved model from:", model_file, "\n")
fit_gram <- readRDS(model_file)
} else {
cat("Fitting model (this may take a while)...\n")
cat("Note: Model fitting requires significant computational resources.\n")
cat("Consider fitting the model separately and saving to fits/fit_gram.rds\n\n")
# Fit with reduced complexity for demonstration
fit_gram <- brm(
correct ~ condition + (1 + condition | subject) + (1 | item),
data = gram_data,
family = bernoulli(),
prior = gram_priors,
chains = 2, # Reduced for faster fitting
iter = 1000,
cores = 2,
backend = "rstan",
refresh = 0
)
# Save for future use
dir.create("fits", showWarnings = FALSE, recursive = TRUE)
saveRDS(fit_gram, model_file)
cat("Model saved to:", model_file, "\n")
}Loading saved model from: fits/fit_gram.rds # Display model summary
cat("\n=== Model Summary ===\n")
=== Model Summary ===print(summary(fit_gram)) Family: bernoulli
Links: mu = logit
Formula: correct ~ condition + (1 + condition | subject) + (1 | item)
Data: gram_data (Number of observations: 2000)
Draws: 2 chains, each with iter = 1000; warmup = 500; thin = 1;
total post-warmup draws = 1000
Multilevel Hyperparameters:
~item (Number of levels: 2)
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sd(Intercept) 0.21 0.24 0.01 0.91 1.03 102 119
~subject (Number of levels: 25)
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS
sd(Intercept) 0.10 0.07 0.00 0.26 1.00 213
sd(conditionB) 0.13 0.09 0.01 0.34 1.00 254
cor(Intercept,conditionB) -0.06 0.44 -0.81 0.76 1.01 427
Tail_ESS
sd(Intercept) 268
sd(conditionB) 560
cor(Intercept,conditionB) 635
Regression Coefficients:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 0.17 0.18 -0.34 0.46 1.02 125 64
conditionB 0.34 0.09 0.17 0.51 1.01 970 657
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).# Bar plot for binary outcomes
pp_check(fit_gram, type = "bars", ndraws = 500) +
ggtitle("Bar plot: Observed vs. Predicted counts")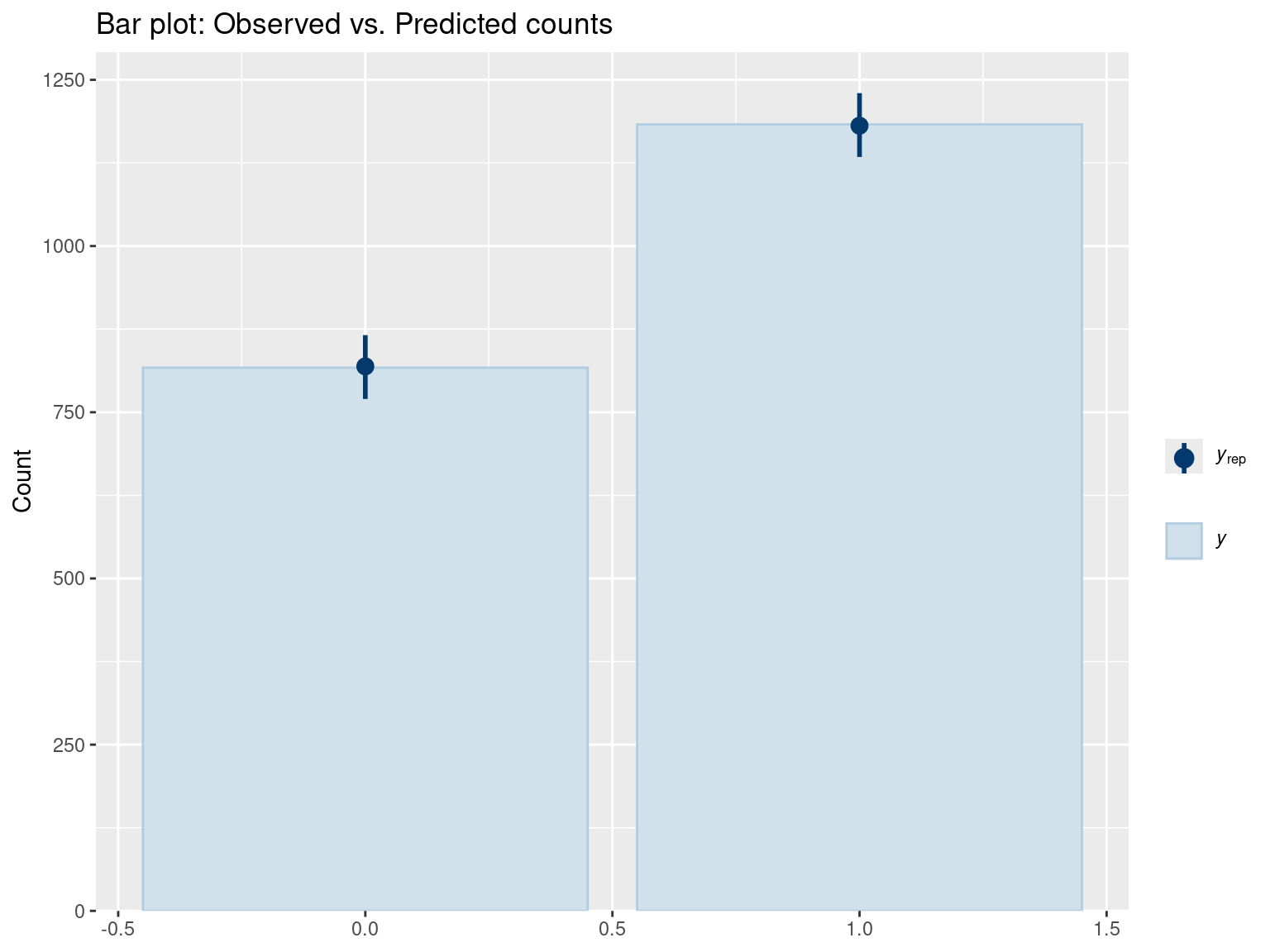
Interpretation:
# Check observed proportion vs. predicted
pp_check(fit_gram, type = "stat", stat = "mean") +
ggtitle("Proportion correct: Observed vs. Predicted")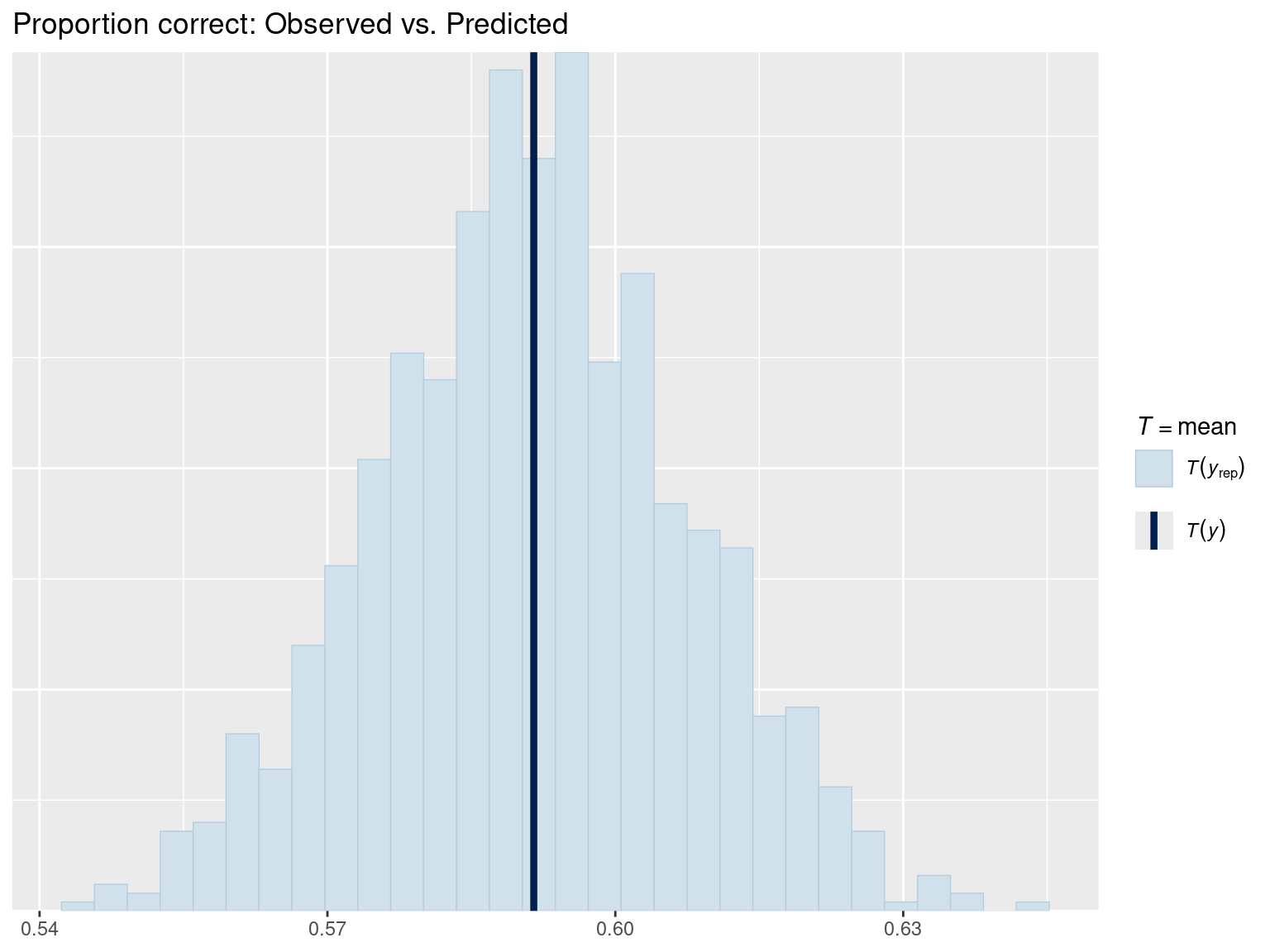
The mean of binary data = proportion of 1’s (proportion correct). This check verifies that the model’s predicted accuracy matches the observed accuracy.
# Error plot for discrete data
pp_check(fit_gram, type = "error_binned") +
ggtitle("Binned error plot")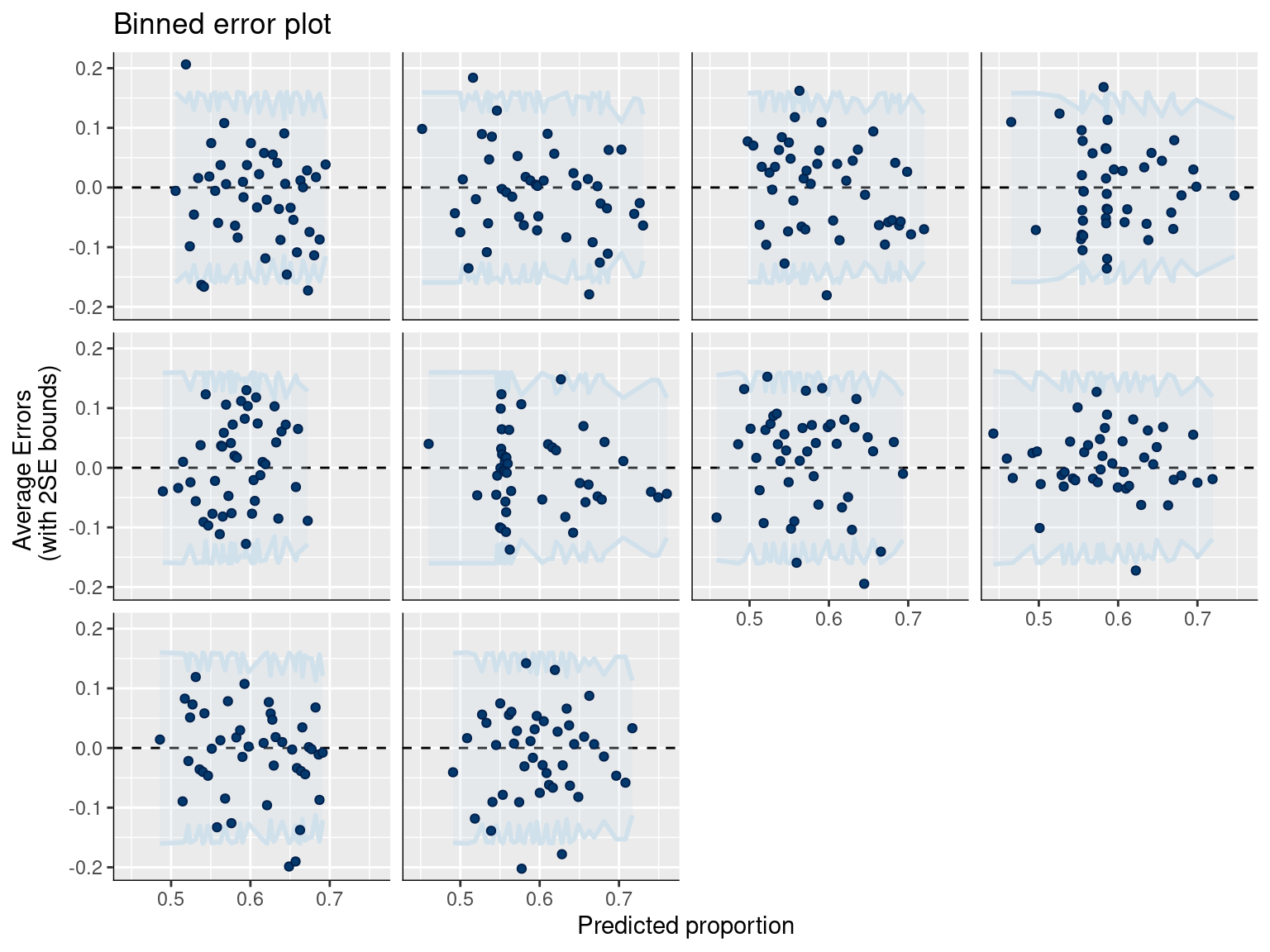
Interpretation:
# Draw from posterior predictive distribution (binary outcomes: 0 or 1)
post_pred <- posterior_predict(fit_gram, ndraws = 1000)
dim(post_pred) # 1000 draws × n observations[1] 1000 2000cat("\nDimensions of posterior predictions:\n")
Dimensions of posterior predictions:cat("Draws:", nrow(post_pred), "\n")Draws: 1000 cat("Observations:", ncol(post_pred), "\n")Observations: 2000 # Calculate overall accuracy
obs_accuracy <- mean(gram_data$correct)
pred_accuracy <- apply(post_pred, 1, mean)
cat("\nOverall Accuracy:\n")
Overall Accuracy:cat("Observed:", round(obs_accuracy, 3), "\n")Observed: 0.601 cat("Predicted (median):", round(median(pred_accuracy), 3), "\n")Predicted (median): 0.591 cat("95% CI:", round(quantile(pred_accuracy, c(0.025, 0.975)), 3), "\n")95% CI: 0.56 0.624 hist(pred_accuracy,
main = "Posterior predictive distribution of accuracy",
xlab = "Proportion correct",
col = "lightblue",
breaks = 30,
xlim = c(0, 1))
abline(v = obs_accuracy, col = "red", lwd = 3, lty = 2)
legend("topleft",
legend = c("Predicted", "Observed"),
col = c("lightblue", "red"),
lwd = c(1, 3),
lty = c(1, 2))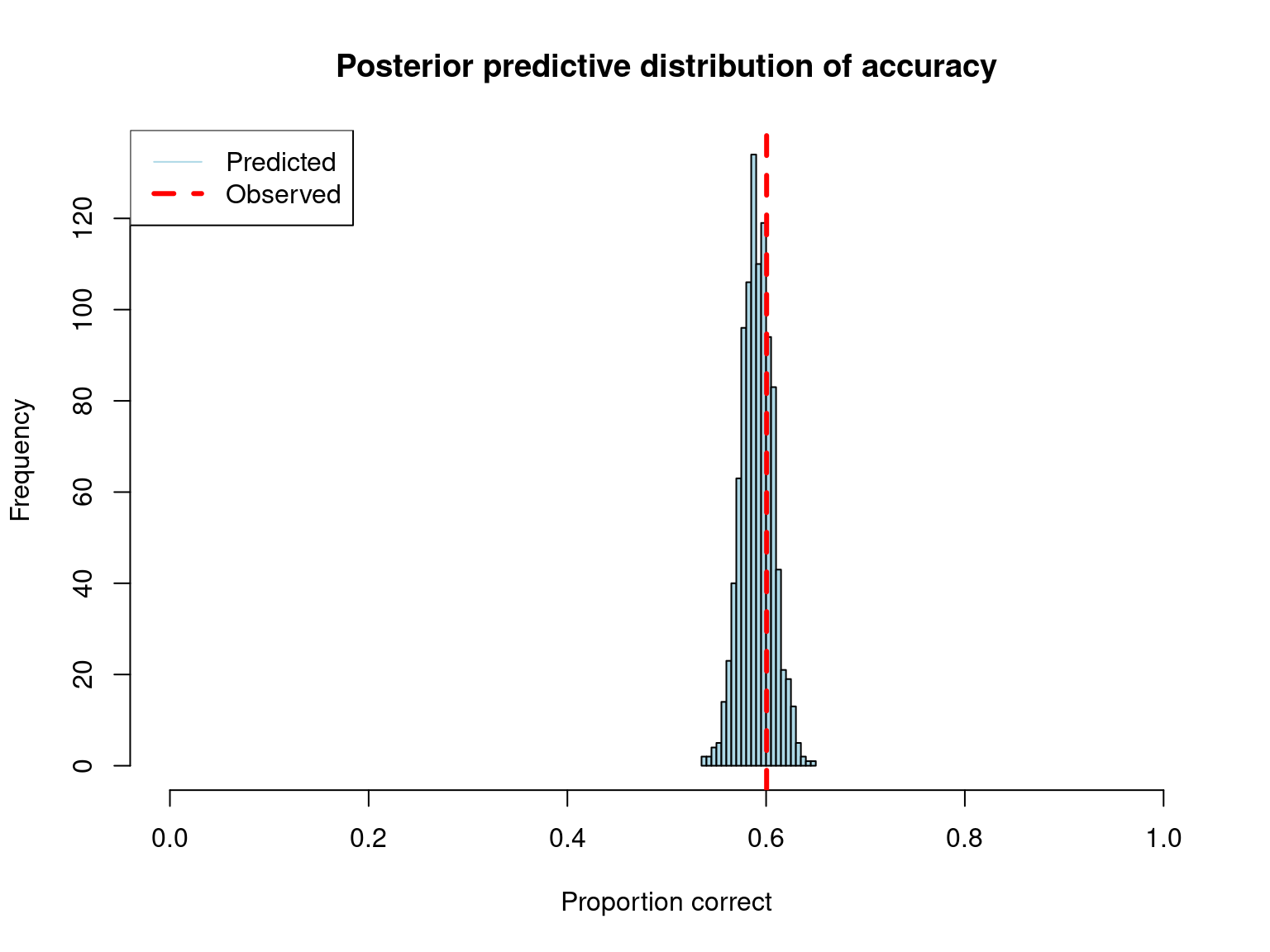
# Check if observed is within reasonable range
if (obs_accuracy < quantile(pred_accuracy, 0.025) ||
obs_accuracy > quantile(pred_accuracy, 0.975)) {
cat("\n⚠ Warning: Observed accuracy outside 95% posterior predictive interval\n")
cat("Model may not be capturing the data well.\n")
} else {
cat("\n✓ Observed accuracy within 95% posterior predictive interval\n")
}
✓ Observed accuracy within 95% posterior predictive interval# Get predicted probabilities (on 0-1 scale)
post_epred <- posterior_epred(fit_gram)
dim(post_epred) # n_draws × n observations[1] 1000 2000cat("\nPosterior expected probabilities:\n")
Posterior expected probabilities:cat("Dimensions:", dim(post_epred), "\n")Dimensions: 1000 2000 # Posterior probability of correct for first observation
cat("\nExample: First observation\n")
Example: First observationcat("Observed outcome:", gram_data$correct[1], "\n")Observed outcome: 0 cat("Predicted P(correct):",
round(quantile(post_epred[, 1], c(0.025, 0.5, 0.975)), 3), "\n")Predicted P(correct): 0.501 0.554 0.614 # Check calibration: do predicted probabilities match observed frequencies?
# Bin by predicted probability and compare to observed proportion
pred_prob_median <- apply(post_epred, 2, median)
# Create bins
n_bins <- 10
bins <- cut(pred_prob_median, breaks = n_bins, include.lowest = TRUE)
calibration_data <- data.frame(
predicted = pred_prob_median,
observed = gram_data$correct,
bin = bins
) %>%
group_by(bin) %>%
summarise(
mean_predicted = mean(predicted),
mean_observed = mean(observed),
n = n()
) %>%
filter(n > 0)
plot(calibration_data$mean_predicted,
calibration_data$mean_observed,
pch = 19,
cex = sqrt(calibration_data$n) / 3,
xlab = "Predicted probability",
ylab = "Observed proportion",
main = "Calibration plot",
xlim = c(0, 1),
ylim = c(0, 1))
abline(0, 1, col = "red", lwd = 2, lty = 2)
grid()
legend("topleft",
legend = "Perfect calibration",
col = "red",
lwd = 2,
lty = 2)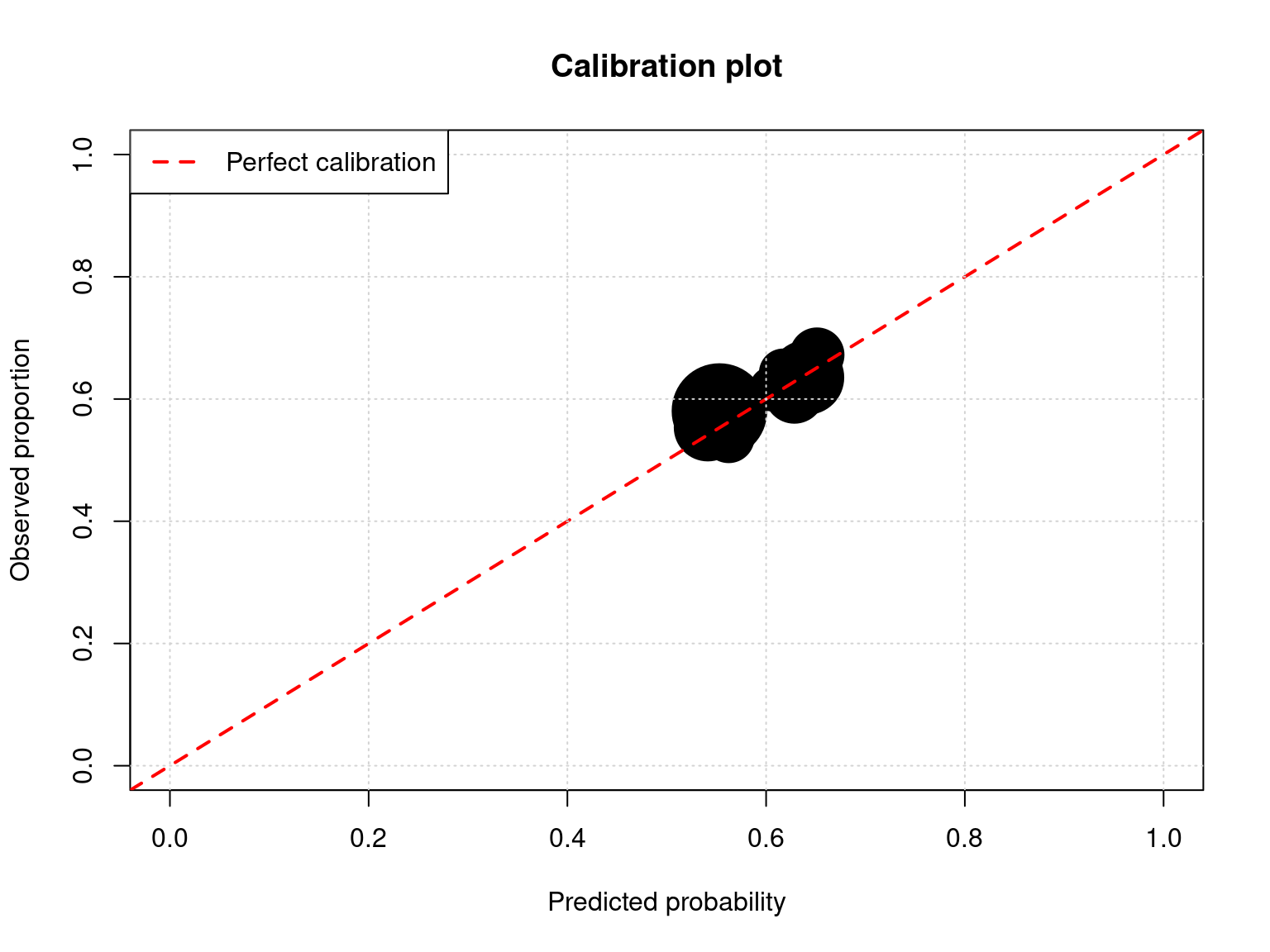
cat("\nCalibration:\n")
Calibration:cat("Points should fall near the diagonal line.\n")Points should fall near the diagonal line.cat("Point size proportional to number of observations.\n")Point size proportional to number of observations.# Observed accuracy by condition
obs_by_condition <- gram_data %>%
group_by(condition) %>%
summarise(
accuracy = mean(correct),
n = n()
)
# Predicted accuracy by condition
gram_data$pred_prob <- apply(post_epred, 2, median)
pred_by_condition <- gram_data %>%
group_by(condition) %>%
summarise(
pred_accuracy = mean(pred_prob)
)
comparison <- left_join(obs_by_condition, pred_by_condition, by = "condition")
cat("\nAccuracy by Condition:\n")
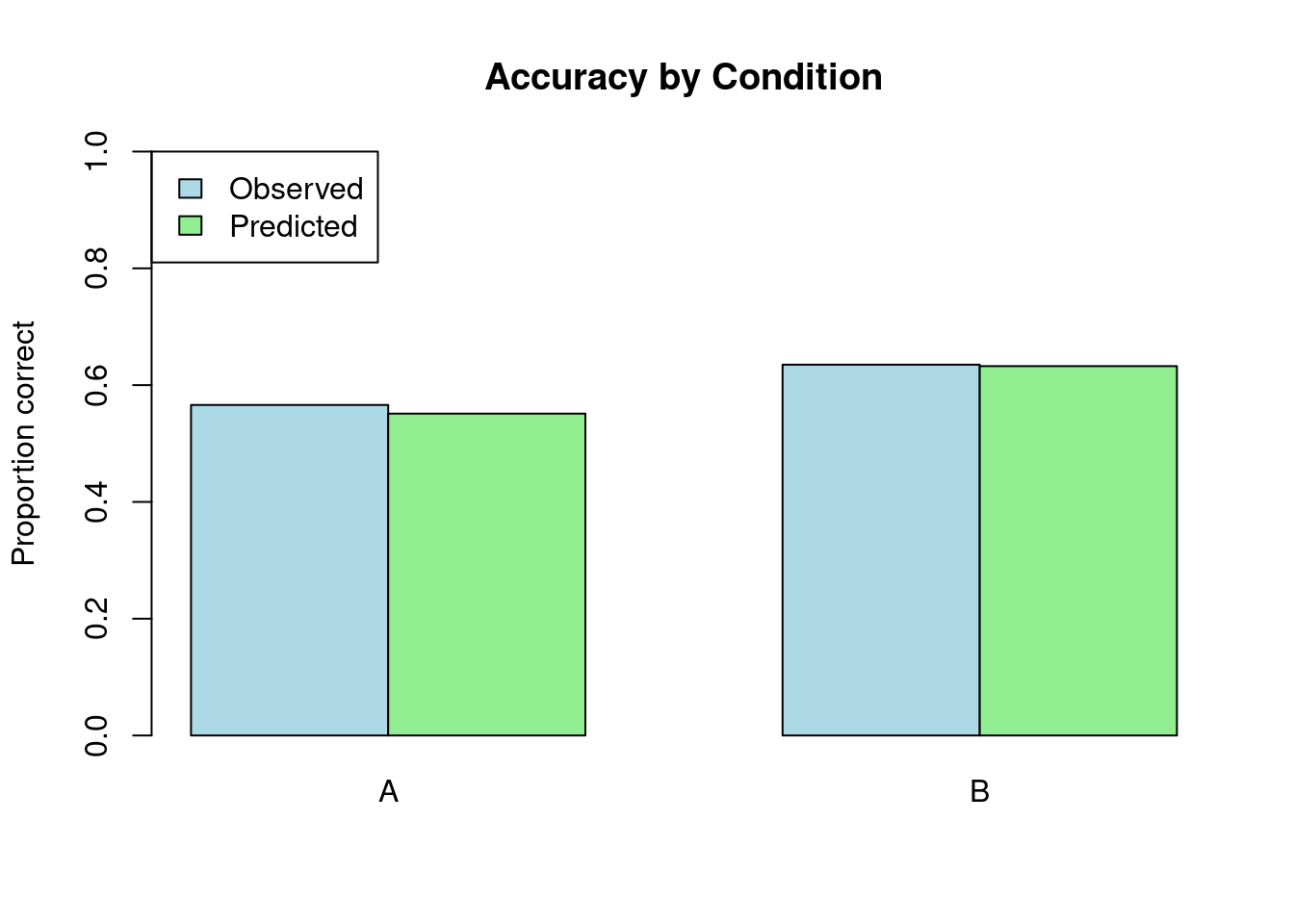
Accuracy by Condition:print(comparison, n = Inf)# A tibble: 2 × 4
condition accuracy n pred_accuracy
<chr> <dbl> <int> <dbl>
1 A 0.566 1000 0.551
2 B 0.635 1000 0.633# Visualize
barplot(
height = rbind(comparison$accuracy, comparison$pred_accuracy),
beside = TRUE,
names.arg = comparison$condition,
col = c("lightblue", "lightgreen"),
main = "Accuracy by Condition",
ylab = "Proportion correct",
ylim = c(0, 1),
legend.text = c("Observed", "Predicted"),
args.legend = list(x = "topleft")
)
# Get predictions for each condition with population-level effects only
new_data <- data.frame(
condition = c("A", "B"),
subject = NA,
item = NA
)
epred_condition <- posterior_epred(fit_gram,
newdata = new_data,
re_formula = NA) # population-level only
# Summarize
condition_summary <- apply(epred_condition, 2, quantile, c(0.025, 0.5, 0.975))
colnames(condition_summary) <- new_data$condition
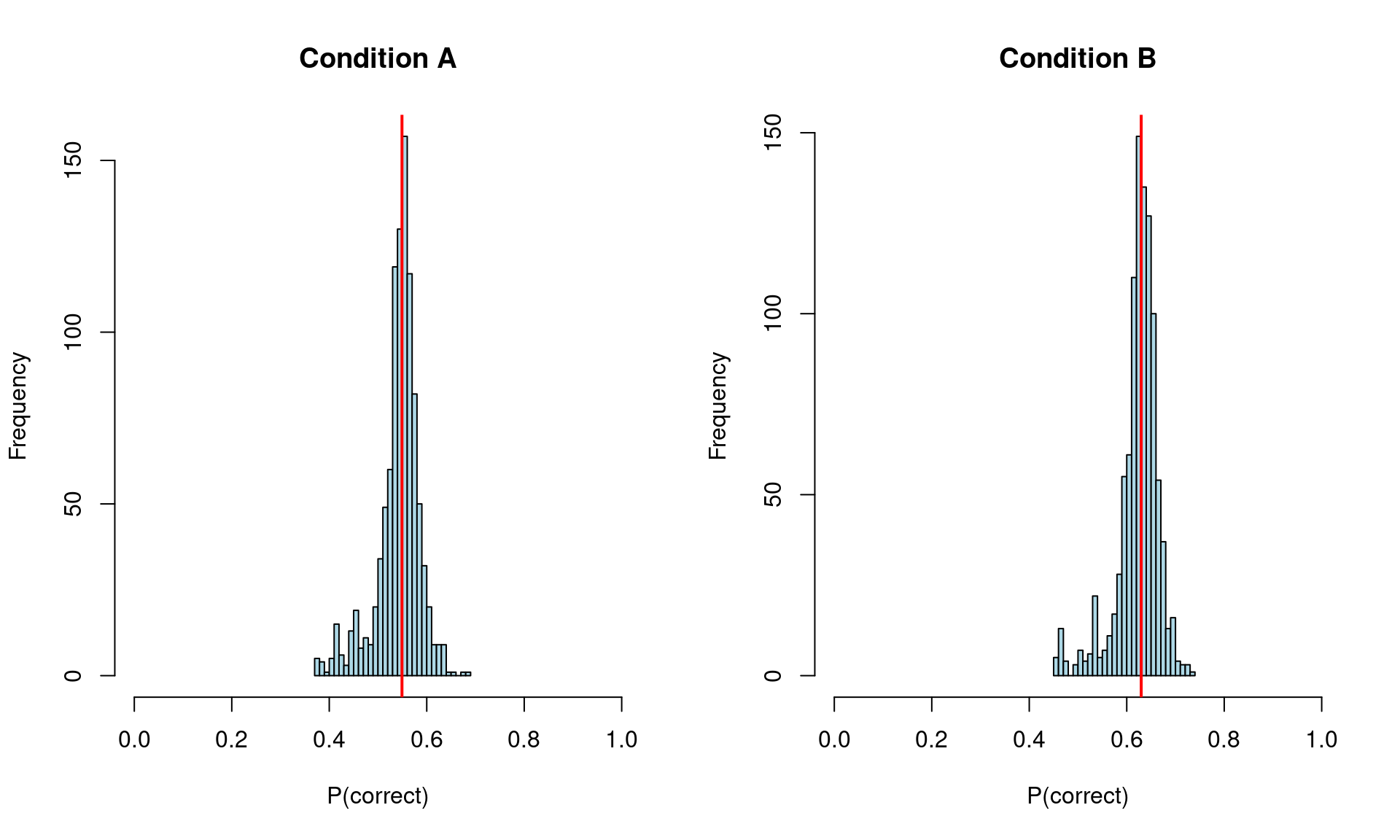
cat("\nPopulation-level P(correct) by condition:\n")
Population-level P(correct) by condition:print(round(condition_summary, 3)) A B
2.5% 0.417 0.501
50% 0.549 0.630
97.5% 0.614 0.690# Plot
par(mfrow = c(1, 2))
for (i in 1:2) {
hist(epred_condition[, i],
main = paste("Condition", new_data$condition[i]),
xlab = "P(correct)",
col = "lightblue",
breaks = 30,
xlim = c(0, 1))
abline(v = median(epred_condition[, i]), col = "red", lwd = 2)
}
| Problem | Diagnosis | Solution |
|---|---|---|
| Poor calibration | Points far from diagonal | Add predictors, check formula |
| Misses condition effects | Different accuracy by condition not captured | Add condition × random effect interaction |
| Predictions too certain | Predicted probs near 0 or 1 | Check priors, may be too strong |
| Predictions too uncertain | Predicted probs all near 0.5 | Add more structure, informative priors |
If posterior predictive checks reveal problems:
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 22.04.5 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.20.so; LAPACK version 3.10.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: Etc/UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] bayesplot_1.14.0 lubridate_1.9.3 forcats_1.0.0 stringr_1.5.1
[5] dplyr_1.1.4 purrr_1.0.2 readr_2.1.5 tidyr_1.3.1
[9] tibble_3.2.1 ggplot2_4.0.0 tidyverse_2.0.0 brms_2.23.0
[13] Rcpp_1.0.13
loaded via a namespace (and not attached):
[1] gtable_0.3.6 tensorA_0.36.2.1 QuickJSR_1.8.1
[4] xfun_0.54 htmlwidgets_1.6.4 inline_0.3.21
[7] lattice_0.22-6 tzdb_0.4.0 vctrs_0.6.5
[10] tools_4.4.1 generics_0.1.3 stats4_4.4.1
[13] parallel_4.4.1 fansi_1.0.6 pkgconfig_2.0.3
[16] Matrix_1.7-0 checkmate_2.3.3 RColorBrewer_1.1-3
[19] S7_0.2.0 distributional_0.5.0 RcppParallel_5.1.11-1
[22] lifecycle_1.0.4 compiler_4.4.1 farver_2.1.2
[25] Brobdingnag_1.2-9 codetools_0.2-20 htmltools_0.5.8.1
[28] yaml_2.3.10 pillar_1.9.0 StanHeaders_2.32.10
[31] bridgesampling_1.1-2 abind_1.4-8 nlme_3.1-164
[34] posterior_1.6.1.9000 rstan_2.32.7 tidyselect_1.2.1
[37] digest_0.6.37 mvtnorm_1.3-3 stringi_1.8.4
[40] reshape2_1.4.4 labeling_0.4.3 fastmap_1.2.0
[43] grid_4.4.1 cli_3.6.5 magrittr_2.0.3
[46] loo_2.8.0 pkgbuild_1.4.8 utf8_1.2.4
[49] withr_3.0.2 scales_1.4.0 backports_1.5.0
[52] estimability_1.5.1 timechange_0.3.0 rmarkdown_2.30
[55] matrixStats_1.5.0 emmeans_2.0.0 gridExtra_2.3
[58] hms_1.1.3 coda_0.19-4.1 evaluate_1.0.1
[61] knitr_1.50 rstantools_2.5.0 rlang_1.1.6
[64] xtable_1.8-4 glue_1.8.0 jsonlite_1.8.9
[67] plyr_1.8.9 R6_2.5.1