Essential Reading
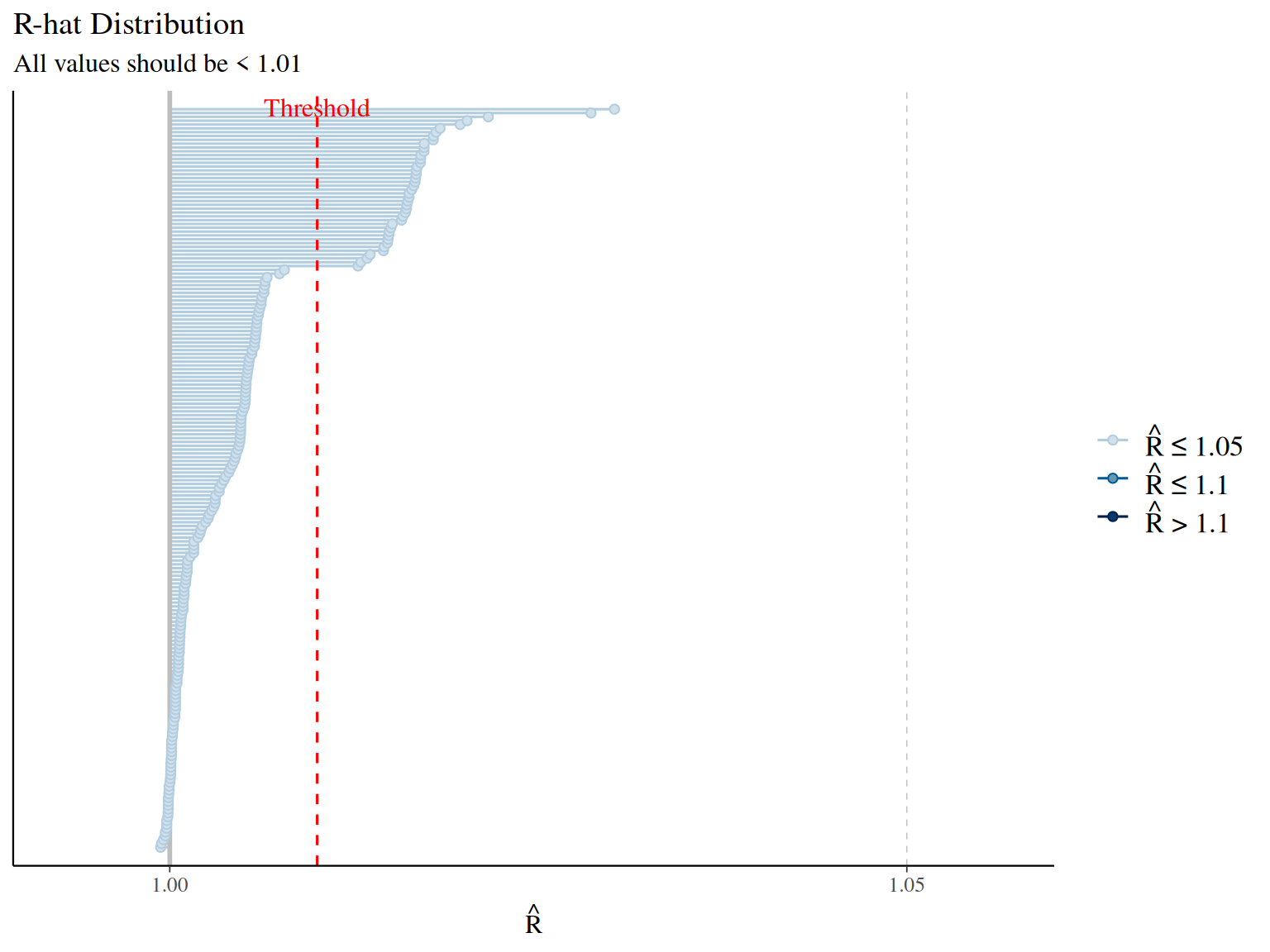
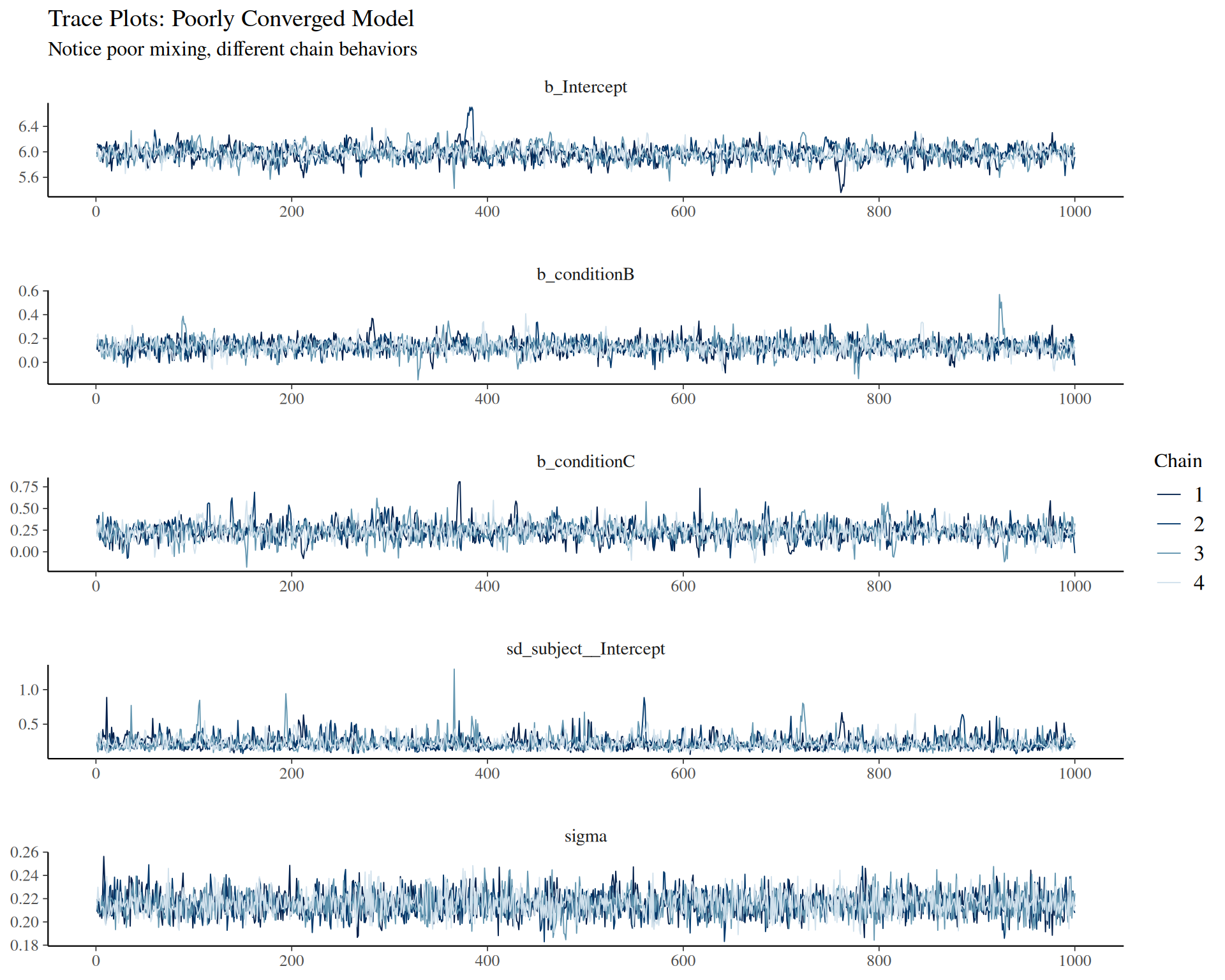
Convergence diagnostics: - Vehtari, A., Gelman, A., Simpson, D., Carpenter, B., & Bürkner, P. C. (2021). Rank-normalization, folding, and localization: An improved R̂ for assessing convergence of MCMC. Bayesian Analysis, 16(2), 667-718. - Modern R-hat diagnostic (current standard)
- Gelman, A., & Rubin, D. B. (1992). Inference from iterative simulation using multiple sequences. Statistical Science, 7(4), 457-472.
- Original R-hat diagnostic
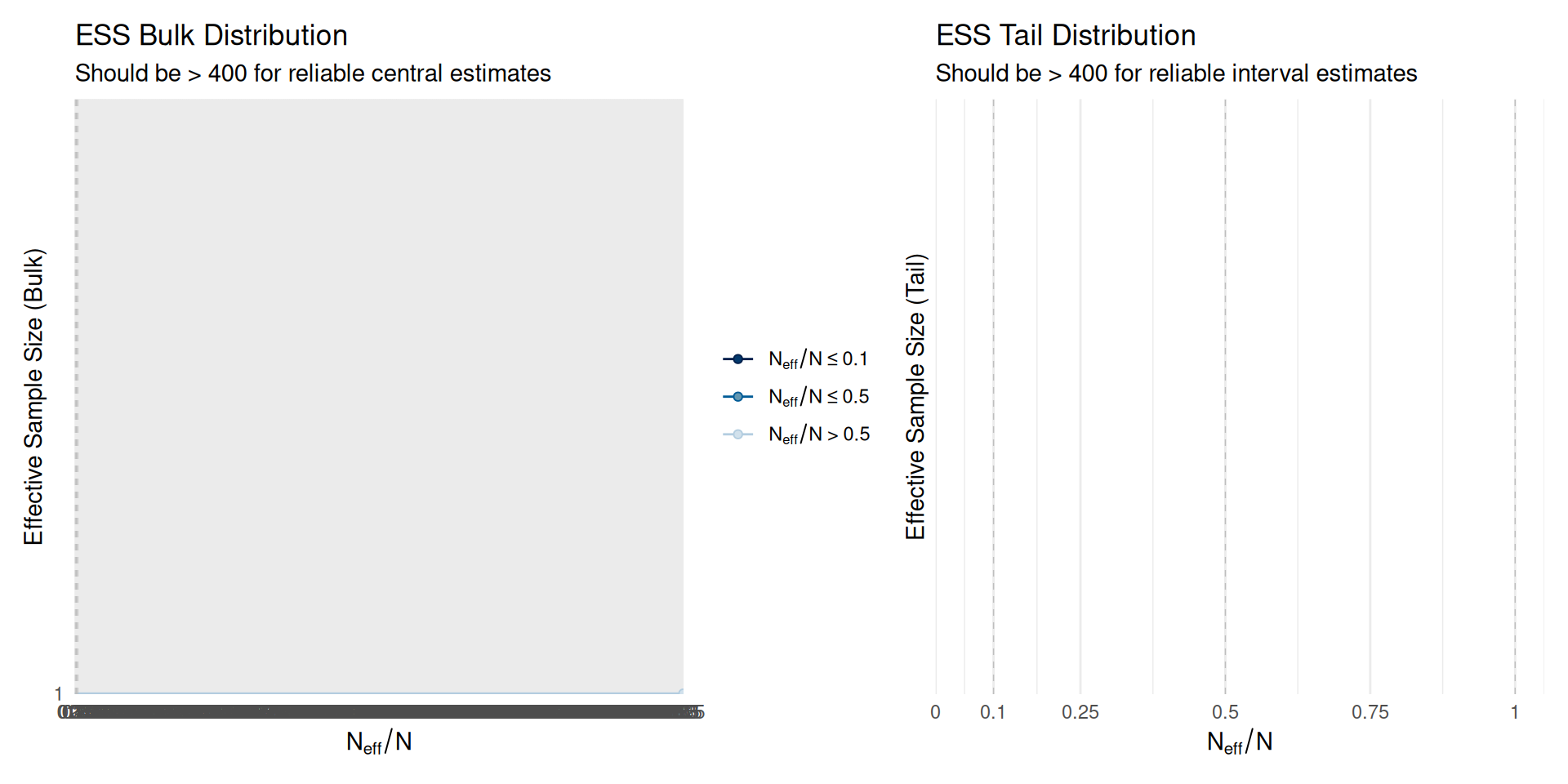
Effective sample size: - Vehtari et al. (2021) [same as above] - ESS bulk and tail diagnostics
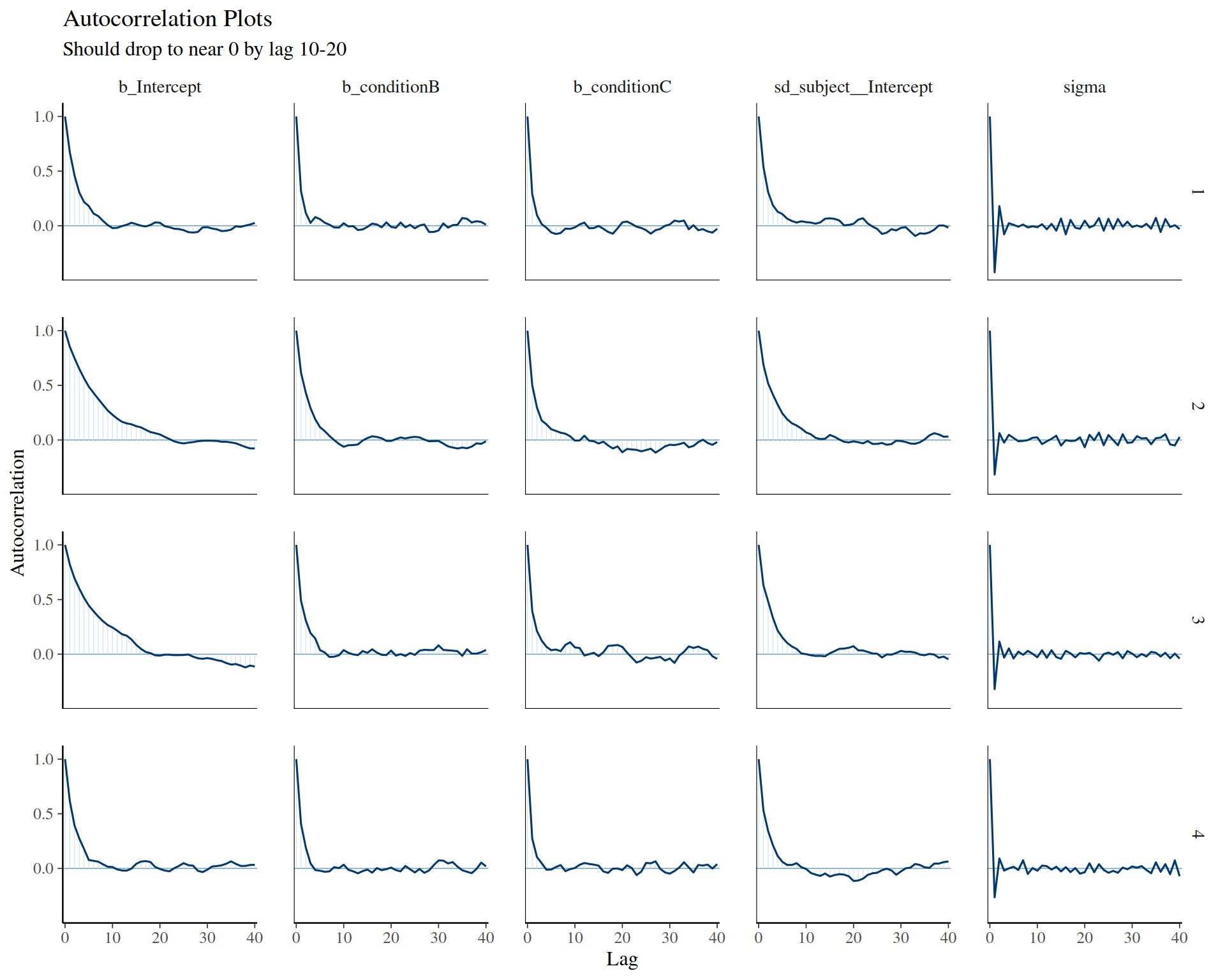
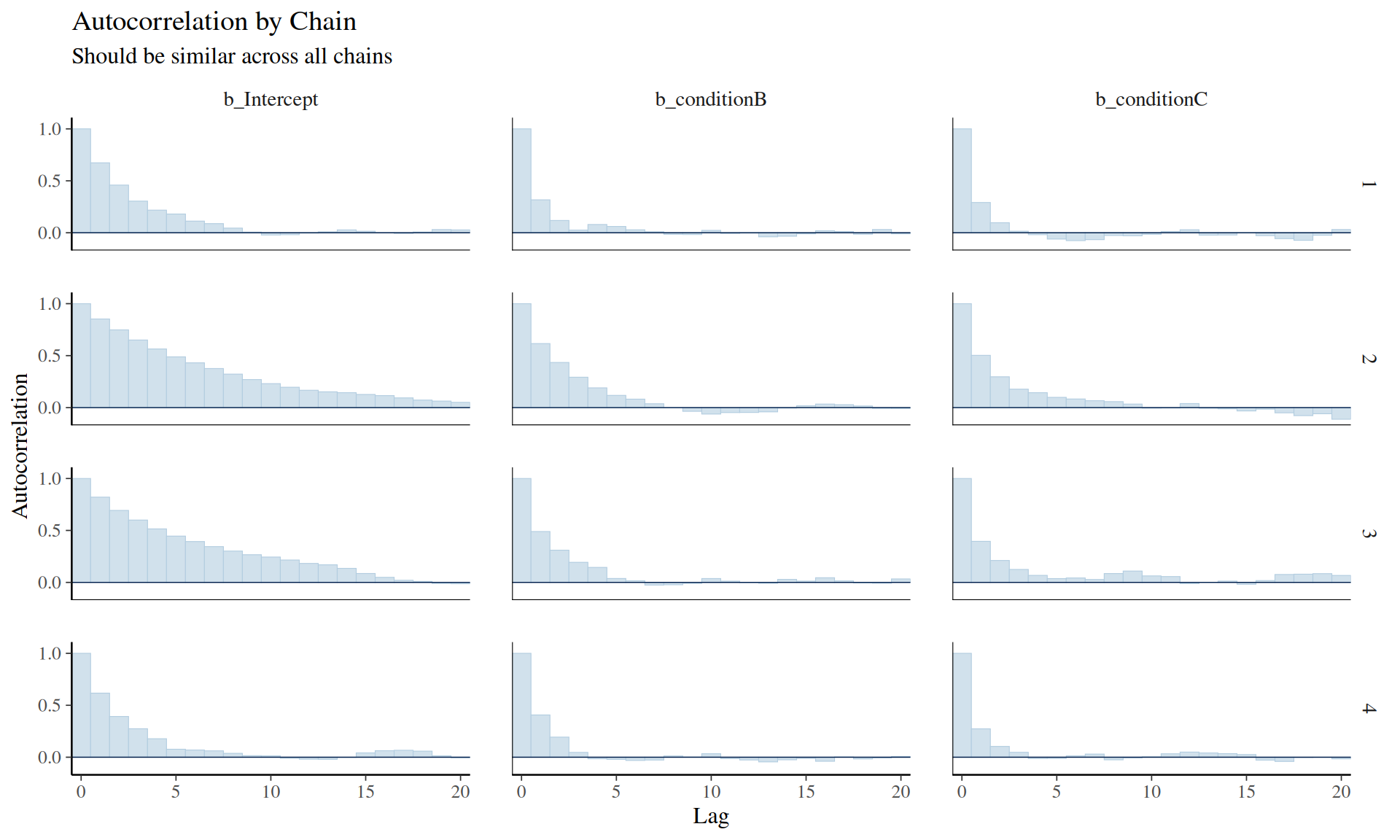
General MCMC diagnostics: - Gelman, A., Carlin, J. B., Stern, H. S., Dunson, D. B., Vehtari, A., & Rubin, D. B. (2013). Bayesian Data Analysis (3rd ed.). CRC Press. - Chapter 11: Basics of Markov Chain Simulation - Gold standard textbook
Practical guidance: - Gabry, J., Simpson, D., Vehtari, A., Betancourt, M., & Gelman, A. (2019). Visualization in Bayesian workflow. Journal of the Royal Statistical Society A, 182(2), 389-402. - Comprehensive workflow including convergence checks