Show code
library(brms)
library(tidyverse)
library(bayesplot)
# Set seed for reproducibility
set.seed(42)Bayesian Mixed Effects Models with brms for Linguists
After fitting your model, validate that it generates data similar to what you observed.
Posterior predictive checks answer: “If I were to generate new data from my fitted model, would it look like my actual data?” This validates that your model has captured the essential structure of your data.
library(brms)
library(tidyverse)
library(bayesplot)
# Set seed for reproducibility
set.seed(42)# Create example RT data (same as in prior checks for consistency)
n_subj <- 20
n_trials <- 50
n_items <- 30
rt_data <- expand.grid(
trial = 1:n_trials,
subject = 1:n_subj,
item = 1:n_items
) %>%
filter(row_number() <= n_subj * n_trials * 3) %>%
mutate(
condition = rep(c("A", "B"), length.out = n()),
# Data generation: two independent noise sources, no random effects
# Total residual SD = sqrt(0.3^2 + 0.1^2) = 0.316
log_rt = rnorm(n(), mean = 6, sd = 0.3) +
(condition == "B") * 0.15 +
rnorm(n(), mean = 0, sd = 0.1),
rt = exp(log_rt)
)
# Define priors
rt_priors <- c(
prior(normal(6, 1.5), class = Intercept),
prior(normal(0, 0.5), class = b),
prior(exponential(1), class = sigma),
prior(exponential(1), class = sd),
prior(lkj(2), class = cor)
)
# Check if model exists, otherwise fit it
model_file <- "fits/fit_rt.rds"
if (file.exists(model_file)) {
cat("Loading saved model from:", model_file, "\n")
fit_rt <- readRDS(model_file)
} else {
cat("Fitting model (this may take a while)...\n")
cat("Note: Model fitting requires significant computational resources.\n")
cat("Consider fitting the model separately and saving to fits/fit_rt.rds\n\n")
# Fit with reduced complexity for demonstration
fit_rt <- brm(
log_rt ~ condition + (1 + condition | subject) + (1 | item),
data = rt_data,
family = gaussian(),
prior = rt_priors,
chains = 2, # Reduced for faster fitting
iter = 1000,
cores = 2,
backend = "rstan",
refresh = 0
)
# Save for future use
dir.create("fits", showWarnings = FALSE, recursive = TRUE)
saveRDS(fit_rt, model_file)
cat("Model saved to:", model_file, "\n")
}Loading saved model from: fits/fit_rt.rds # Display model summary
cat("\n=== Model Summary ===\n")
=== Model Summary ===print(summary(fit_rt)) Family: gaussian
Links: mu = identity
Formula: log_rt ~ condition + (1 + condition | subject) + (1 | item)
Data: rt_data (Number of observations: 3000)
Draws: 2 chains, each with iter = 1000; warmup = 500; thin = 1;
total post-warmup draws = 1000
Multilevel Hyperparameters:
~item (Number of levels: 3)
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sd(Intercept) 0.02 0.02 0.00 0.07 1.01 392 323
~subject (Number of levels: 20)
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS
sd(Intercept) 0.02 0.01 0.00 0.04 1.00 432
sd(conditionB) 0.02 0.01 0.00 0.06 1.00 343
cor(Intercept,conditionB) -0.07 0.46 -0.82 0.83 1.01 513
Tail_ESS
sd(Intercept) 528
sd(conditionB) 461
cor(Intercept,conditionB) 591
Regression Coefficients:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 5.99 0.01 5.96 6.02 1.00 700 436
conditionB 0.16 0.01 0.13 0.19 1.00 1339 699
Further Distributional Parameters:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sigma 0.32 0.00 0.31 0.33 1.01 2149 698
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).# Default: density overlay of observed vs. simulated data
pp_check(fit_rt, ndraws = 100) +
ggtitle("Density overlay: Observed vs. Posterior predictions")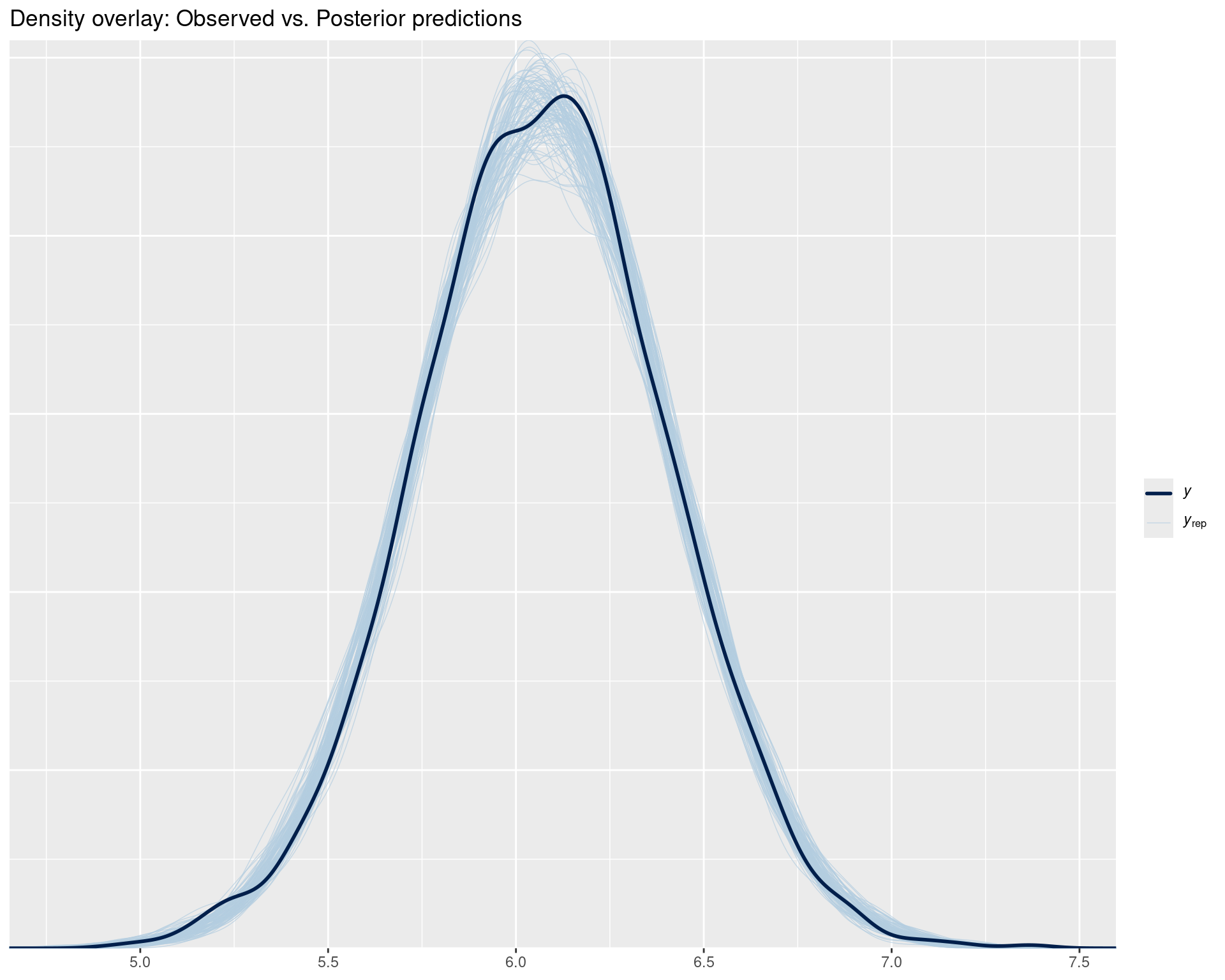
Interpretation:
y = observed data) should be among the blue lines (yrep = posterior predictions)# Did we get the mean right?
p1 <- pp_check(fit_rt, type = "stat", stat = "mean") +
ggtitle("Mean: Observed vs. Predicted")
# Did we get the spread right?
p2 <- pp_check(fit_rt, type = "stat", stat = "sd") +
ggtitle("SD: Observed vs. Predicted")
# Extreme values?
p3 <- pp_check(fit_rt, type = "stat", stat = "min") +
ggtitle("Minimum: Observed vs. Predicted")
p4 <- pp_check(fit_rt, type = "stat", stat = "max") +
ggtitle("Maximum: Observed vs. Predicted")
# Display all plots
library(patchwork)
(p1 | p2) / (p3 | p4)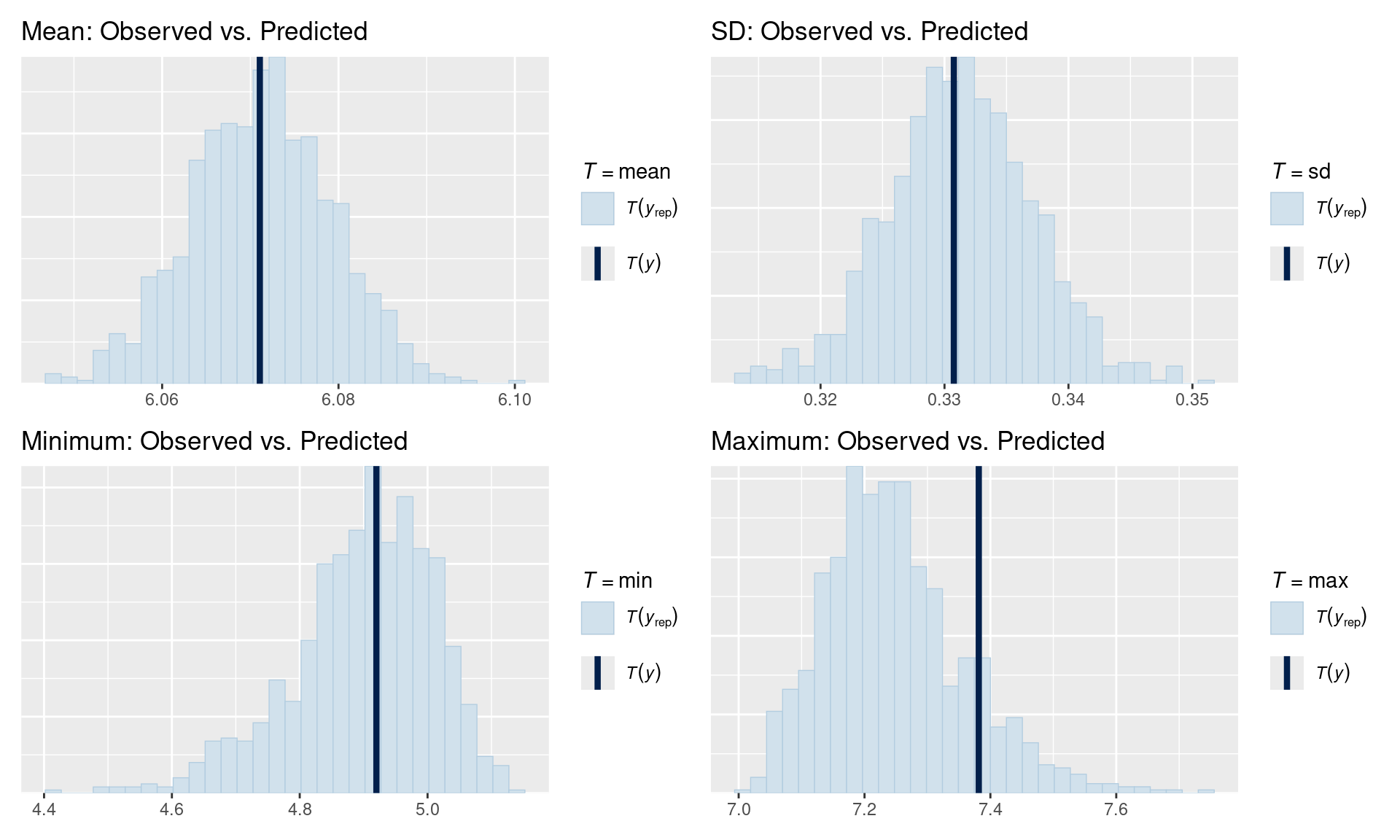
# Draw from posterior predictive distribution
post_pred <- posterior_predict(fit_rt, ndraws = 1000)
dim(post_pred) # 1000 draws × n observations[1] 1000 3000cat("\nDimensions of posterior predictions:\n")
Dimensions of posterior predictions:cat("Draws:", nrow(post_pred), "\n")Draws: 1000 cat("Observations:", ncol(post_pred), "\n")Observations: 3000 # Compare observed vs. predicted means
obs_mean <- mean(rt_data$log_rt)
pred_mean <- apply(post_pred, 1, mean)
hist(pred_mean,
main = "Posterior predictive distribution of mean log-RT",
xlab = "Mean log-RT",
col = "lightblue",
breaks = 30)
abline(v = obs_mean, col = "red", lwd = 3, lty = 2)
legend("topright",
legend = c("Predicted", "Observed"),
col = c("lightblue", "red"),
lwd = c(1, 3),
lty = c(1, 2))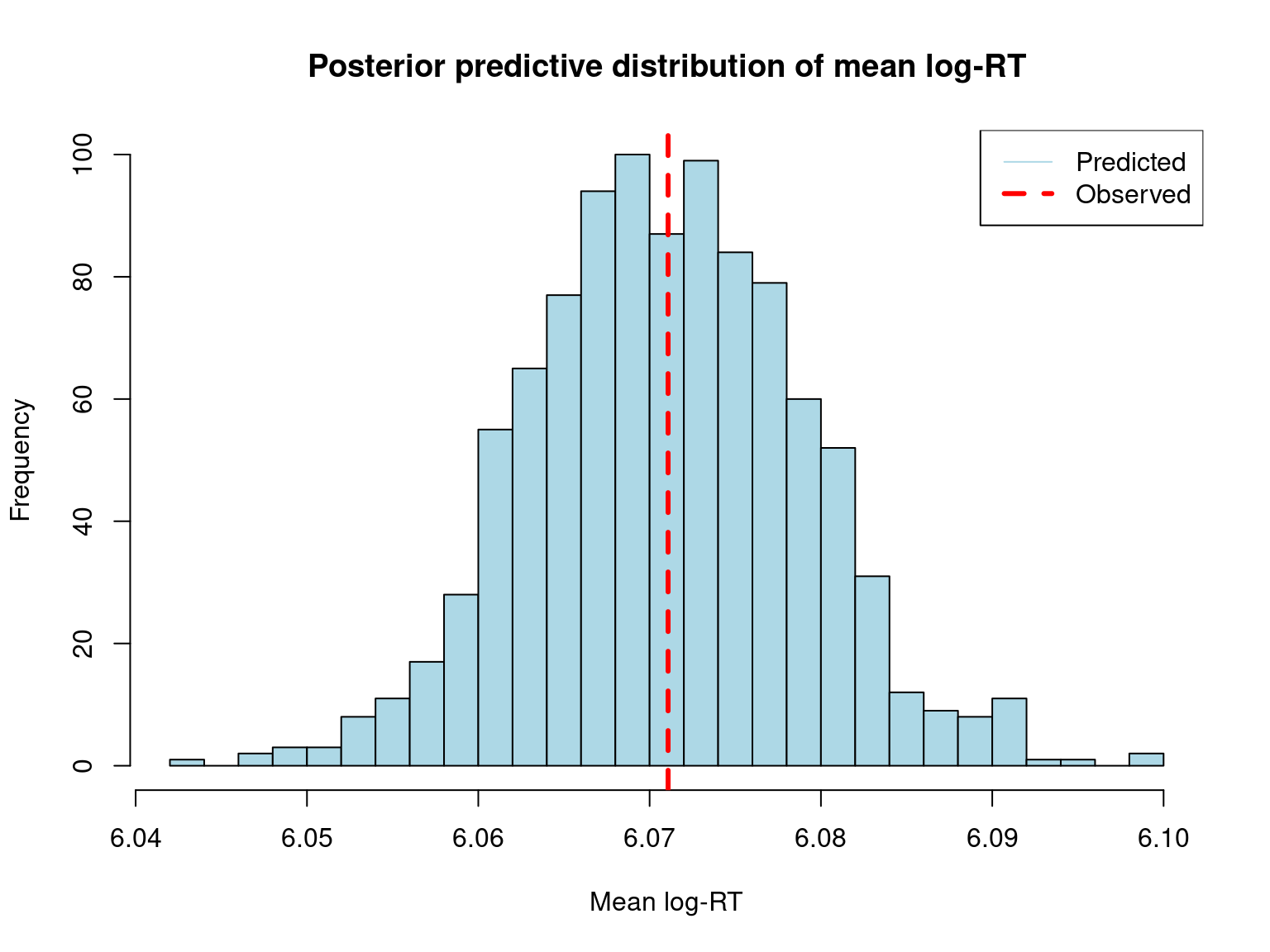
cat("\nObserved mean log-RT:", round(obs_mean, 3), "\n")
Observed mean log-RT: 6.071 cat("Predicted mean log-RT (median):", round(median(pred_mean), 3), "\n")Predicted mean log-RT (median): 6.071 cat("95% CI for predicted mean:",
round(quantile(pred_mean, c(0.025, 0.975)), 3), "\n")95% CI for predicted mean: 6.056 6.087 # Check 95% posterior predictive interval
post_pred_interval <- apply(post_pred, 2, quantile, c(0.025, 0.975))
# Roughly 95% of observed values should fall within their interval
coverage <- mean(rt_data$log_rt > post_pred_interval[1,] &
rt_data$log_rt < post_pred_interval[2,])
cat("\nPosterior Predictive Interval Coverage:\n")
Posterior Predictive Interval Coverage:cat("Proportion of observations within 95% interval:", round(coverage, 3), "\n")Proportion of observations within 95% interval: 0.951 cat("Expected: ~0.95\n")Expected: ~0.95if (coverage < 0.90) {
cat("\n⚠ Warning: Coverage is lower than expected.\n")
cat("Model may be overconfident or missing important structure.\n")
} else if (coverage > 0.98) {
cat("\n⚠ Warning: Coverage is higher than expected.\n")
cat("Model may be too uncertain or overfitted.\n")
} else {
cat("\n✓ Coverage looks good!\n")
}
✓ Coverage looks good!# Group predictions by condition
rt_data_cond <- rt_data %>%
group_by(condition) %>%
summarise(
obs_mean = mean(log_rt),
obs_sd = sd(log_rt)
)
# Get posterior predictions by condition
post_pred_A <- posterior_predict(fit_rt,
newdata = filter(rt_data, condition == "A"),
ndraws = 1000)
post_pred_B <- posterior_predict(fit_rt,
newdata = filter(rt_data, condition == "B"),
ndraws = 1000)
pred_mean_A <- apply(post_pred_A, 1, mean)
pred_mean_B <- apply(post_pred_B, 1, mean)
# Plot comparison
par(mfrow = c(1, 2))
hist(pred_mean_A,
main = "Condition A: Mean log-RT",
xlab = "Mean log-RT",
col = "lightblue",
breaks = 30,
xlim = range(c(pred_mean_A, pred_mean_B)))
abline(v = rt_data_cond$obs_mean[1], col = "red", lwd = 3, lty = 2)
hist(pred_mean_B,
main = "Condition B: Mean log-RT",
xlab = "Mean log-RT",
col = "lightgreen",
breaks = 30,
xlim = range(c(pred_mean_A, pred_mean_B)))
abline(v = rt_data_cond$obs_mean[2], col = "red", lwd = 3, lty = 2)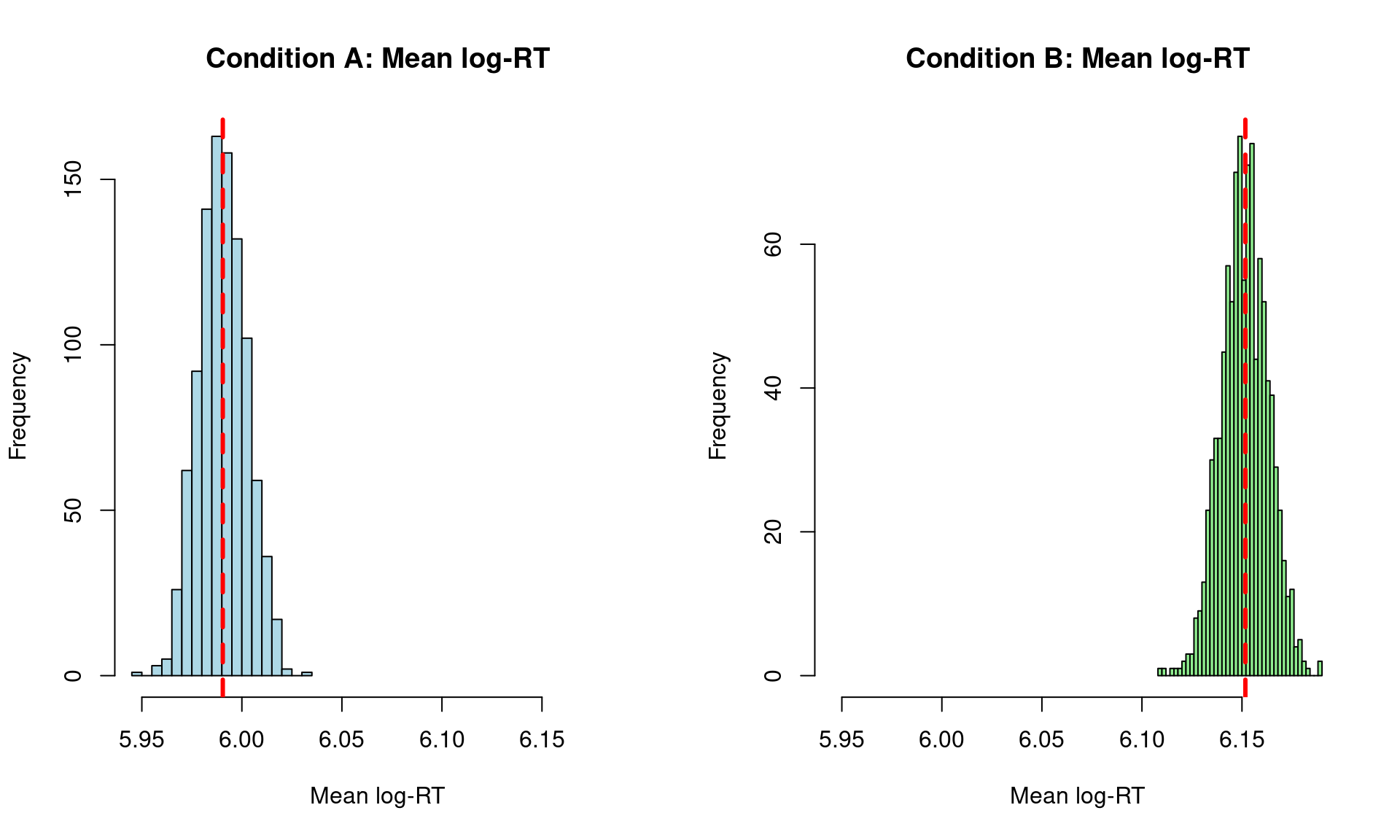
cat("\nBy Condition:\n")
By Condition:cat("Condition A - Observed:", round(rt_data_cond$obs_mean[1], 3), "\n")Condition A - Observed: 5.99 cat("Condition A - Predicted:", round(median(pred_mean_A), 3), "\n")Condition A - Predicted: 5.99 cat("Condition B - Observed:", round(rt_data_cond$obs_mean[2], 3), "\n")Condition B - Observed: 6.152 cat("Condition B - Predicted:", round(median(pred_mean_B), 3), "\n")Condition B - Predicted: 6.151 This section demonstrates how the posterior predictive distribution converges to the true data-generating value as we add more observations. We’ll fit models with increasing amounts of data and examine how well each recovers the residual SD.
# Sample sizes to test
n_obs_levels <- c(0, 1, 2, 3, 4, 5, 10, 50, 100, 1000, 3000)
# Function to fit model with subset of data
fit_with_n_obs <- function(n_obs, full_data, priors) {
if (n_obs == 0) {
# Prior predictive only
sampled_data <- full_data[1:100, ] # Need some data structure
fit <- brm(
log_rt ~ condition + (1 + condition | subject) + (1 | item),
data = sampled_data,
family = gaussian(),
prior = priors,
chains = 2,
iter = 1000,
cores = 2,
sample_prior = "only",
backend = "cmdstanr",
refresh = 0,
silent = 2
)
} else {
# For very small samples, use intercept-only model
if (n_obs <= 5) {
# Just grab first n observations
sampled_data <- full_data %>%
slice_head(n = n_obs)
# Intercept-only model (no predictors)
formula <- log_rt ~ 1
# Simplified priors for intercept-only
model_priors <- c(
prior(normal(6, 1.5), class = Intercept),
prior(exponential(1), class = sigma)
)
} else if (n_obs <= 20) {
# Stratified sampling for small samples
sampled_data <- full_data %>%
group_by(condition) %>%
slice_sample(n = max(2, n_obs %/% 2)) %>%
ungroup() %>%
slice_head(n = n_obs)
# Simple fixed effects only
formula <- log_rt ~ condition
# Priors for fixed effects model
model_priors <- c(
prior(normal(6, 1.5), class = Intercept),
prior(normal(0, 0.5), class = b),
prior(exponential(1), class = sigma)
)
} else if (n_obs <= 100) {
# Random sampling with simple random effects
sampled_data <- full_data %>%
slice_sample(n = min(n_obs, nrow(full_data)), replace = FALSE)
# Add subject random intercepts only
formula <- log_rt ~ condition + (1 | subject)
model_priors <- c(
prior(normal(6, 1.5), class = Intercept),
prior(normal(0, 0.5), class = b),
prior(exponential(1), class = sigma),
prior(exponential(1), class = sd)
)
} else {
# Full model for larger samples
sampled_data <- full_data %>%
slice_sample(n = min(n_obs, nrow(full_data)), replace = FALSE)
formula <- log_rt ~ condition + (1 + condition | subject) + (1 | item)
model_priors <- priors
}
fit <- brm(
formula = formula,
data = sampled_data,
family = gaussian(),
prior = model_priors,
chains = 2,
iter = 1000,
cores = 2,
backend = "cmdstanr",
refresh = 0,
silent = 2
)
}
return(fit)
}
# Fit models (this may take several minutes)
cat("Fitting models with varying sample sizes...\n")Fitting models with varying sample sizes...cat("This will take a few minutes. Progress:\n")This will take a few minutes. Progress:fits_list <- list()
for (i in seq_along(n_obs_levels)) {
n <- n_obs_levels[i]
cat(sprintf(" [%2d/11] n = %4d observations...", i, n))
# Check for cached model
cache_file <- sprintf("fits/fit_rt_n%04d.rds", n)
if (file.exists(cache_file)) {
fits_list[[i]] <- readRDS(cache_file)
cat(" (loaded from cache)\n")
} else {
fits_list[[i]] <- fit_with_n_obs(n, rt_data, rt_priors)
# Save to cache
dir.create("fits", showWarnings = FALSE, recursive = TRUE)
saveRDS(fits_list[[i]], cache_file)
cat(" (fitted and cached)\n")
}
} [ 1/11] n = 0 observations... (loaded from cache)
[ 2/11] n = 1 observations... (loaded from cache)
[ 3/11] n = 2 observations... (loaded from cache)
[ 4/11] n = 3 observations... (loaded from cache)
[ 5/11] n = 4 observations... (loaded from cache)
[ 6/11] n = 5 observations... (loaded from cache)
[ 7/11] n = 10 observations... (loaded from cache)
[ 8/11] n = 50 observations... (loaded from cache)
[ 9/11] n = 100 observations... (loaded from cache)
[10/11] n = 1000 observations... (loaded from cache)
[11/11] n = 3000 observations... (loaded from cache)cat("\nGenerating posterior predictive checks...\n")
Generating posterior predictive checks...# Create comparison plots
library(patchwork)
plots_list <- lapply(seq_along(n_obs_levels), function(i) {
n <- n_obs_levels[i]
fit <- fits_list[[i]]
# Generate pp_check for SD
# Use "ppd" for prior predictive (n=0), "ppc" for posterior predictive (n>0)
p <- pp_check(fit, type = "stat", stat = "sd", prefix = if(n == 0) "ppd" else "ppc") +
ggtitle(sprintf("n = %d", n)) +
theme_minimal() +
theme(
plot.title = element_text(size = 10, face = "bold"),
axis.title = element_text(size = 8),
axis.text = element_text(size = 7)
)
# Add true value line and sampling uncertainty band
if (n > 0) {
# True total SD from data generation includes:
# 1. Residual variance: sqrt(0.3^2 + 0.1^2) = 0.316
# 2. Fixed effect variance from balanced condition: sqrt(0.5 * 0.5 * 0.15^2) = 0.075
# Total SD = sqrt(0.3^2 + 0.1^2 + 0.5 * 0.5 * 0.15^2) ≈ 0.325
true_sigma <- sqrt(0.3^2 + 0.1^2 + 0.5 * 0.5 * 0.15^2)
p <- p + geom_vline(xintercept = true_sigma, color = "red", linetype = "dashed", linewidth = 1)
# Add 95% CI for where T(y) would fall when sampling from true model
# For sample SD: Use chi-squared distribution
# Sample variance s^2 ~ sigma^2 * chi-squared(n-1) / (n-1)
# Therefore sample SD follows scaled chi distribution
df <- n - 1
# 95% CI for sample SD using chi-squared quantiles
lower_band <- true_sigma * sqrt(qchisq(0.025, df) / df)
upper_band <- true_sigma * sqrt(qchisq(0.975, df) / df)
# Add shaded region showing where single observed SD typically falls
p <- p + annotate("rect",
xmin = lower_band,
xmax = upper_band,
ymin = -Inf, ymax = Inf,
alpha = 0.15, fill = "red")
}
return(p)
})
# Arrange in grid
wrap_plots(plots_list, ncol = 4) +
plot_annotation(
title = "Convergence of Posterior Predictions for Residual SD",
subtitle = "Red dashed line shows true data-generating SD = 0.3 | Prior: exponential(1)",
theme = theme(
plot.title = element_text(size = 14, face = "bold"),
plot.subtitle = element_text(size = 11)
)
)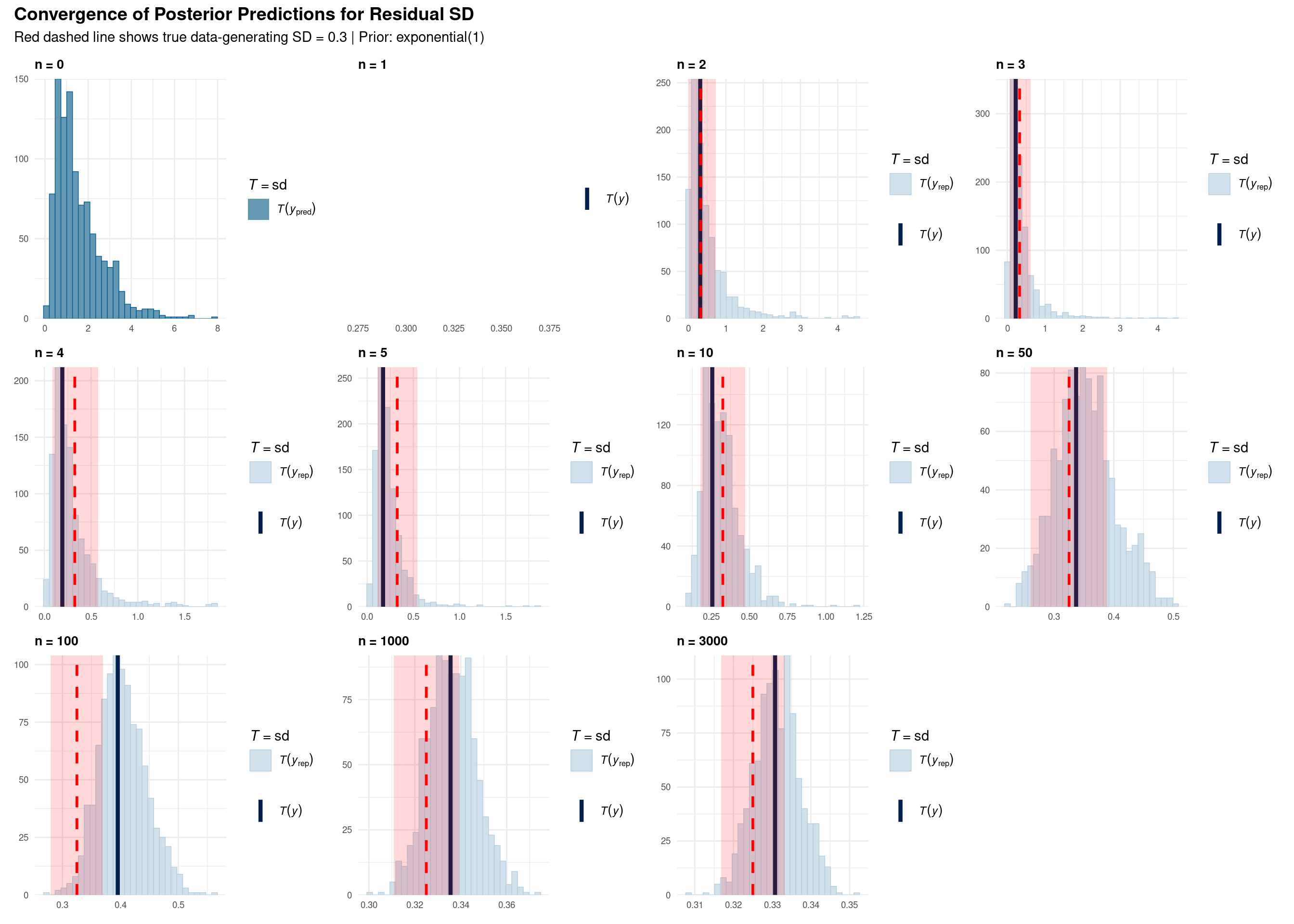
Each subplot shows a posterior predictive check for the standard deviation statistic:
Visual Elements:
What convergence looks like:
Key observations from the progression:
n = 0 (Prior only): Wide uncertainty, no learning from data yet. Prior allows SD anywhere from 0 to ~3.
n = 1-5 (Very few observations): Posterior still heavily influenced by prior. High uncertainty remains. T(y) can be far from truth.
n = 10-50 (Small samples): Beginning to converge toward true value (0.3), but still substantial uncertainty. Histogram narrowing.
n = 100-1000 (Medium samples): Clear convergence visible. Posterior concentrates around 0.3. T(y) aligns with T(y_pred).
n = 3000 (Full data): Tight posterior distribution centered on true value. Prior influence minimal. Very narrow histogram.
This demonstrates:
An important cautionary case: n = 100
Notice at n = 100, T(y) falls outside the red sampling bands. This is a Type I error scenario (occurring ~5% of the time by chance). What are the consequences?
Key insight: Even when the model “correctly” learns from the data (blue histogram centered on T(y)), if the observed data arose from sampling variability (T(y) outside red bands), downstream inferences about effect sizes and standardized parameters will be systematically biased. As sample size increases (n = 1000, 3000), the probability of such extreme sampling variation diminishes, and estimates converge to truth.
# Extract sigma estimates from each model
sigma_summary <- map_dfr(seq_along(n_obs_levels), function(i) {
n <- n_obs_levels[i]
fit <- fits_list[[i]]
# Extract sigma posterior
sigma_draws <- as_draws_df(fit) %>% pull(sigma)
tibble(
n_obs = n,
median = median(sigma_draws),
lower = quantile(sigma_draws, 0.025),
upper = quantile(sigma_draws, 0.975),
width = upper - lower
)
})
# Plot convergence
ggplot(sigma_summary, aes(x = n_obs, y = median)) +
geom_line(linewidth = 1) +
geom_ribbon(aes(ymin = lower, ymax = upper), alpha = 0.2) +
geom_hline(yintercept = 0.3, color = "red", linetype = "dashed", linewidth = 1) +
geom_point(size = 3) +
scale_x_log10(breaks = n_obs_levels) +
labs(
title = "Convergence of Residual SD Estimate",
subtitle = "Red line: true value (0.3) | Ribbon: 95% credible interval",
x = "Number of Observations (log scale)",
y = "Estimated Residual SD (sigma)"
) +
theme_minimal()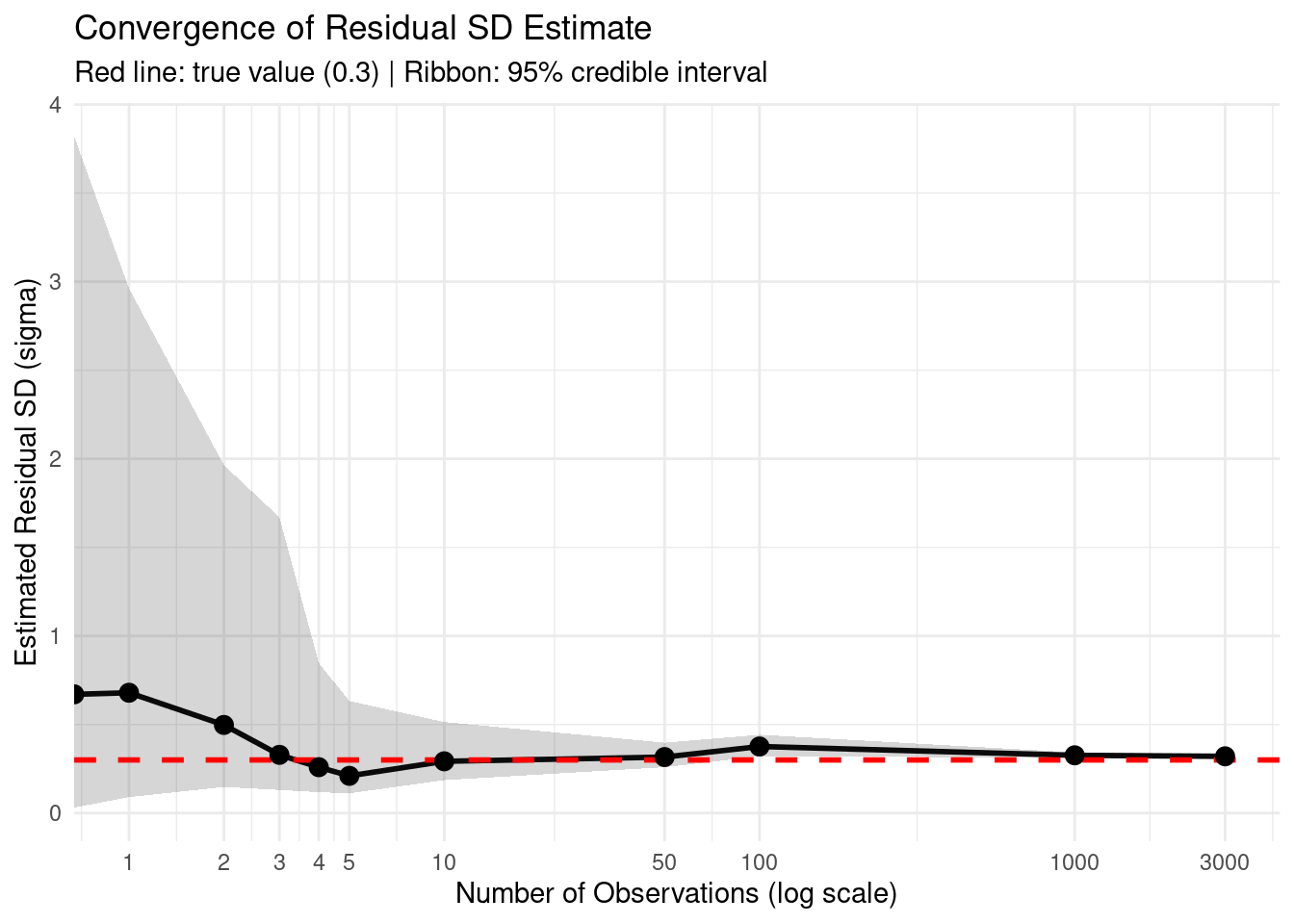
# Print summary table
cat("\n=== Convergence Summary ===\n\n")
=== Convergence Summary ===print(sigma_summary, n = Inf)# A tibble: 11 × 5
n_obs median lower upper width
<dbl> <dbl> <dbl> <dbl> <dbl>
1 0 0.670 0.0310 3.82 3.79
2 1 0.679 0.0902 2.96 2.87
3 2 0.498 0.148 1.96 1.82
4 3 0.329 0.131 1.67 1.54
5 4 0.259 0.118 0.847 0.730
6 5 0.210 0.112 0.632 0.521
7 10 0.292 0.186 0.514 0.328
8 50 0.316 0.259 0.397 0.138
9 100 0.376 0.320 0.442 0.122
10 1000 0.326 0.313 0.343 0.0294
11 3000 0.320 0.313 0.328 0.0152Practical takeaway: With the exponential(1) prior and ~100 observations, the posterior estimate becomes quite accurate. The wide Intercept prior normal(6, 1.5) doesn’t prevent learning the correct residual SD because they model different aspects of the data.
Now let’s examine how the population mean estimate converges with increasing data:
# Create comparison plots for MEAN
plots_list_mean <- lapply(seq_along(n_obs_levels), function(i) {
n <- n_obs_levels[i]
fit <- fits_list[[i]]
# Generate pp_check for MEAN
p <- pp_check(fit, type = "stat", stat = "mean", prefix = if(n == 0) "ppd" else "ppc") +
ggtitle(sprintf("n = %d", n)) +
theme_minimal() +
theme(
plot.title = element_text(size = 10, face = "bold"),
axis.title = element_text(size = 8),
axis.text = element_text(size = 7)
)
# Add true value line and sampling uncertainty band
if (n > 0) {
# True population mean: 6 + 0.15/2 (balanced A/B, B is 0.15 higher)
true_mean <- 6 + 0.15 / 2
p <- p + geom_vline(xintercept = true_mean, color = "red", linetype = "dashed", linewidth = 1)
# Add sampling distribution around true value
# For mean: SE = population_sd / sqrt(n)
# True total SD includes residual variance + fixed effect variance
true_sigma <- sqrt(0.3^2 + 0.1^2 + 0.5 * 0.5 * 0.15^2)
se_of_mean <- true_sigma / sqrt(n)
lower_band <- true_mean - 1.96 * se_of_mean
upper_band <- true_mean + 1.96 * se_of_mean
# Add shaded region showing expected sampling variation
p <- p + annotate("rect",
xmin = lower_band,
xmax = upper_band,
ymin = -Inf, ymax = Inf,
alpha = 0.1, fill = "red")
}
return(p)
})
# Arrange in grid
wrap_plots(plots_list_mean, ncol = 4) +
plot_annotation(
title = "Convergence of Posterior Predictions for Population Mean",
subtitle = "Red dashed line shows true data-generating mean = 6 (log-RT scale) | Prior: normal(6, 1.5)",
theme = theme(
plot.title = element_text(size = 14, face = "bold"),
plot.subtitle = element_text(size = 11)
)
)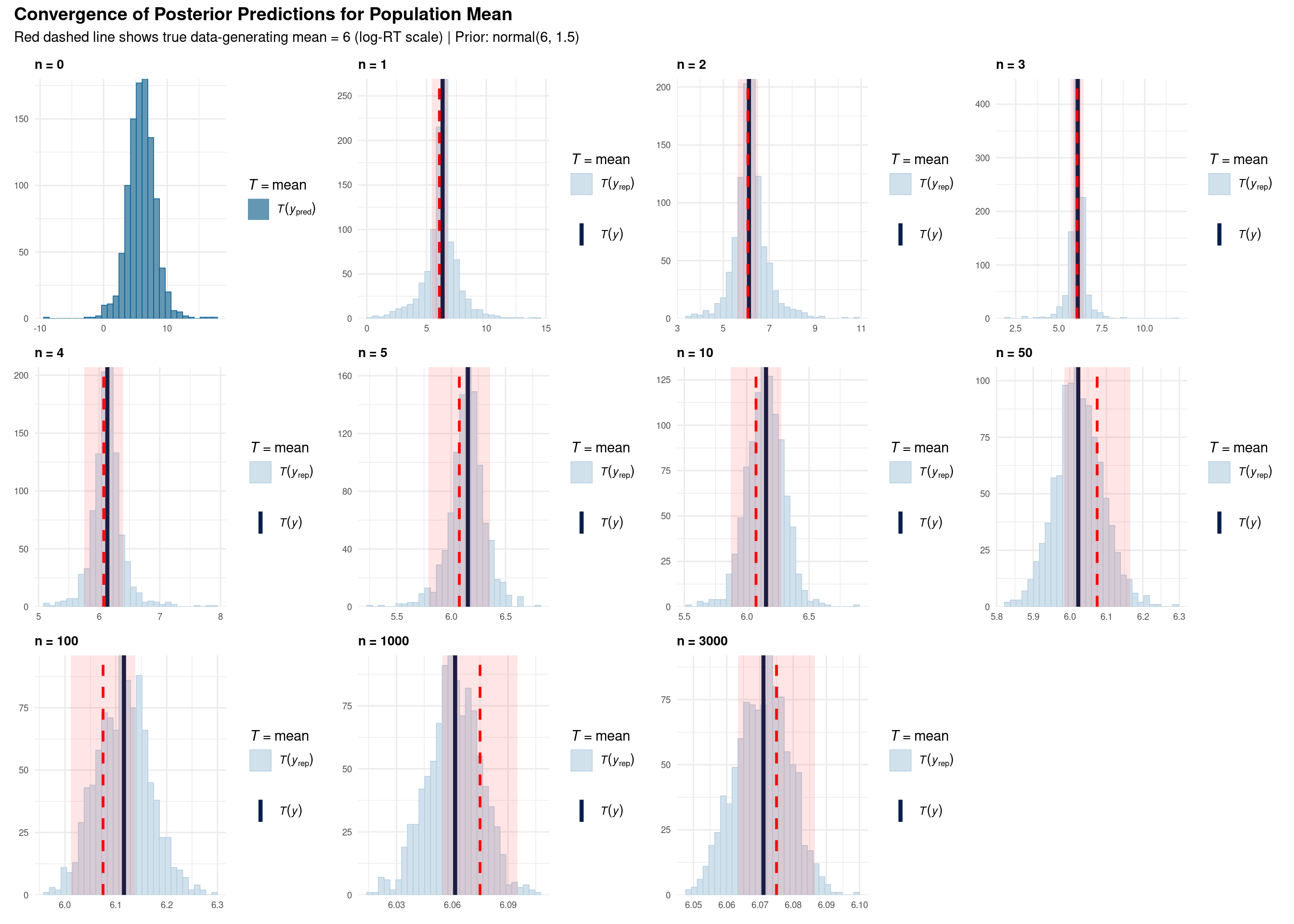
Each subplot shows a posterior predictive check for the mean statistic:
Visual Elements:
Comparison with SD plots:
# Extract Intercept (mean) estimates from each model
mean_summary <- map_dfr(seq_along(n_obs_levels), function(i) {
n <- n_obs_levels[i]
fit <- fits_list[[i]]
# Extract Intercept posterior (represents population mean in intercept-only models)
intercept_draws <- as_draws_df(fit) %>% pull(b_Intercept)
tibble(
n_obs = n,
median = median(intercept_draws),
lower = quantile(intercept_draws, 0.025),
upper = quantile(intercept_draws, 0.975),
width = upper - lower
)
})
# Plot convergence
ggplot(mean_summary, aes(x = n_obs, y = median)) +
geom_line(linewidth = 1) +
geom_ribbon(aes(ymin = lower, ymax = upper), alpha = 0.2) +
geom_hline(yintercept = 6, color = "red", linetype = "dashed", linewidth = 1) +
geom_point(size = 3) +
scale_x_log10(breaks = n_obs_levels) +
labs(
title = "Convergence of Population Mean Estimate (Intercept)",
subtitle = "Red line: true value (6.0) | Ribbon: 95% credible interval",
x = "Number of Observations (log scale)",
y = "Estimated Mean (log-RT scale)"
) +
theme_minimal()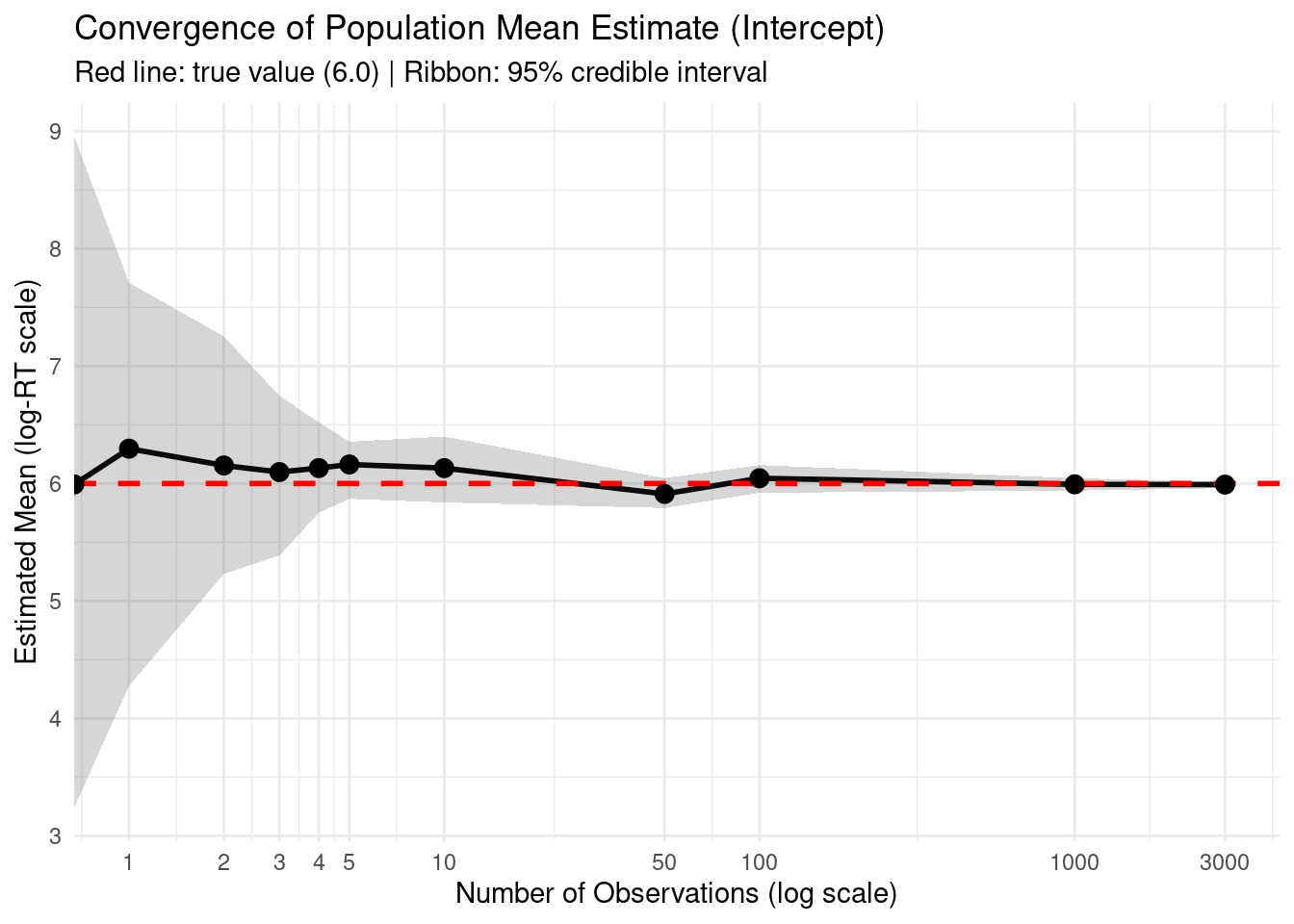
# Print summary table
cat("\n=== Mean Convergence Summary ===\n\n")
=== Mean Convergence Summary ===print(mean_summary, n = Inf)# A tibble: 11 × 5
n_obs median lower upper width
<dbl> <dbl> <dbl> <dbl> <dbl>
1 0 5.99 3.24 8.96 5.72
2 1 6.30 4.28 7.71 3.43
3 2 6.15 5.23 7.25 2.02
4 3 6.10 5.39 6.75 1.36
5 4 6.13 5.75 6.52 0.769
6 5 6.16 5.87 6.36 0.486
7 10 6.13 5.84 6.40 0.561
8 50 5.91 5.79 6.05 0.254
9 100 6.04 5.92 6.16 0.236
10 1000 5.99 5.94 6.04 0.105
11 3000 5.99 5.96 6.02 0.0546cat("\n=== Comparison: Mean vs. SD Convergence ===\n\n")
=== Comparison: Mean vs. SD Convergence ===cat("Credible interval width at n=10:\n")Credible interval width at n=10:cat(" Mean:", round(mean_summary$width[mean_summary$n_obs == 10], 3), "\n") Mean: 0.561 cat(" SD: ", round(sigma_summary$width[sigma_summary$n_obs == 10], 3), "\n\n") SD: 0.328 cat("Credible interval width at n=100:\n")Credible interval width at n=100:cat(" Mean:", round(mean_summary$width[mean_summary$n_obs == 100], 3), "\n") Mean: 0.236 cat(" SD: ", round(sigma_summary$width[sigma_summary$n_obs == 100], 3), "\n\n") SD: 0.122 cat("→ Mean converges faster: CI narrows more quickly with increasing data\n")→ Mean converges faster: CI narrows more quickly with increasing dataThis analysis demonstrates the original question: Why does normal(6, 1.5) for the Intercept not prevent learning sigma = 0.3?
They model different aspects:
normal(6, 1.5): Uncertainty about the population mean
exponential(1): Uncertainty about residual variation
With sufficient data (n > 100):
| Problem | Diagnosis | Solution |
|---|---|---|
| Model predictions too narrow | SD of posterior predictions < SD of data | Relax priors, check formula |
| Model predictions too wide | SD of posterior predictions >> SD of data | Tighten priors, add more structure |
| Misses condition effects | Mean differs dramatically by condition | Add condition × random effect interaction |
| Extreme value mismatch | Min/max far from observed | Check for outliers, consider robust models |
If posterior predictive checks reveal problems:
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 22.04.5 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.20.so; LAPACK version 3.10.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: Etc/UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] patchwork_1.3.2 bayesplot_1.14.0 lubridate_1.9.3 forcats_1.0.0
[5] stringr_1.5.1 dplyr_1.1.4 purrr_1.0.2 readr_2.1.5
[9] tidyr_1.3.1 tibble_3.2.1 ggplot2_4.0.0 tidyverse_2.0.0
[13] brms_2.23.0 Rcpp_1.0.13
loaded via a namespace (and not attached):
[1] gtable_0.3.6 tensorA_0.36.2.1 QuickJSR_1.8.1
[4] xfun_0.54 htmlwidgets_1.6.4 processx_3.8.4
[7] inline_0.3.21 lattice_0.22-6 tzdb_0.4.0
[10] ps_1.8.1 vctrs_0.6.5 tools_4.4.1
[13] generics_0.1.3 stats4_4.4.1 parallel_4.4.1
[16] fansi_1.0.6 cmdstanr_0.9.0 pkgconfig_2.0.3
[19] Matrix_1.7-0 checkmate_2.3.3 RColorBrewer_1.1-3
[22] S7_0.2.0 distributional_0.5.0 RcppParallel_5.1.11-1
[25] lifecycle_1.0.4 compiler_4.4.1 farver_2.1.2
[28] Brobdingnag_1.2-9 codetools_0.2-20 htmltools_0.5.8.1
[31] yaml_2.3.10 pillar_1.9.0 StanHeaders_2.32.10
[34] bridgesampling_1.1-2 abind_1.4-8 nlme_3.1-164
[37] posterior_1.6.1.9000 rstan_2.32.7 tidyselect_1.2.1
[40] digest_0.6.37 mvtnorm_1.3-3 stringi_1.8.4
[43] reshape2_1.4.4 labeling_0.4.3 fastmap_1.2.0
[46] grid_4.4.1 cli_3.6.5 magrittr_2.0.3
[49] loo_2.8.0 pkgbuild_1.4.8 utf8_1.2.4
[52] withr_3.0.2 scales_1.4.0 backports_1.5.0
[55] estimability_1.5.1 timechange_0.3.0 rmarkdown_2.30
[58] matrixStats_1.5.0 emmeans_2.0.0 gridExtra_2.3
[61] hms_1.1.3 coda_0.19-4.1 evaluate_1.0.1
[64] knitr_1.50 rstantools_2.5.0 rlang_1.1.6
[67] xtable_1.8-4 glue_1.8.0 jsonlite_1.8.9
[70] plyr_1.8.9 R6_2.5.1